Big geodata management and analysis using GRASS GIS
FOSS4G 2018 workshop
Veronica Andreo, Marco Ciolli, Luca Delucchi and Markus Neteler
Course outline:
- Introduction to GRASS GIS
- Introduction to GRASS Temporal Framework
- Hands-on to raster time series processing
Software requirements:
- GRASS GIS 7.4: download | OSGeo-live
- GRASS GIS Add-ons: i.modis.download, i.modis.import, i.sentinel.download, i.sentinel.import, i.sentinel.preproc, i.sentinel.mask, r.in.srtm.region and v.strds.stats
- the following Python libraries: pyModis, sentinelsat, pandas, ply
Sample data:
- Main dataset: North Carolina basic location (50 MB)
- extra "mapset"
modis_lst: monthly land surface temperature (LST) from MODIS sensor (MOD11B3.006) for North Carolina (2015-2016). This dataset is required if you cannot download MODIS data from NASA
- extra "mapset"
GRASS GIS introduction
Working with GRASS GIS is not much different from any other GIS. Just a few commonly used terms need to be introduced first.GRASS DATABASE, LOCATION and MAPSET
The GRASS DATABASE (also called "GISDBASE") is an existing directory which contains all GRASS GIS projects. These projects are organized in subdirectories called LOCATIONs.A LOCATION is defined by its coordinate system, map projection and geographical boundaries.
MAPSETs are subdirectories within Locations. In a MAPSET you can organize GIS maps thematically, geographically, by project or however you prefer.
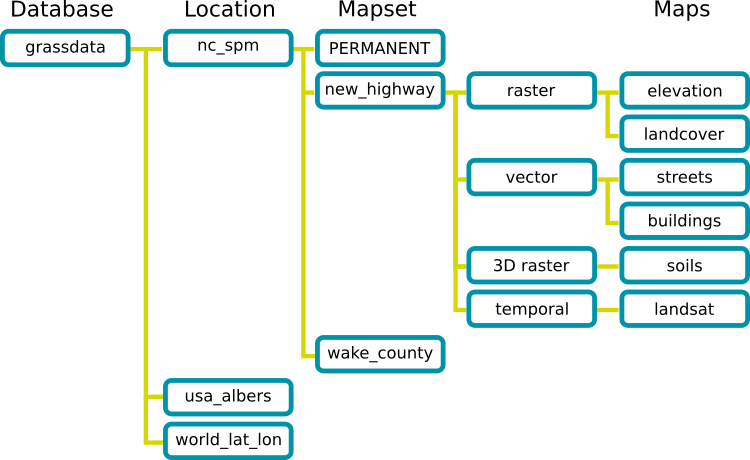
GRASS DATABASE, LOCATIONs and MAPSETs
Raster and vector maps
GRASS GIS is able to read most GIS data formats directly (mainly done through the GDAL library). GRASS GIS has its own internal formats to manage raster and vector data, so your data have to be imported or linked into a GRASS LOCATION/MAPSET.The GRASS GIS vector format is topological, this means that adjacent geographic components in a single vector map are related to each other. For example, in a non-topological GIS if two areas share a common border that border would be digitized twice and also stored in a duplicate manner. In a topological GIS such as GRASS GIS, this border exists only once and it is shared between these two areas. The topological representation of vector data helps to produce and maintain vector maps with clean geometry. Moreover, it enables certain analyses that can not be conducted with non-topological or spaghetti data.
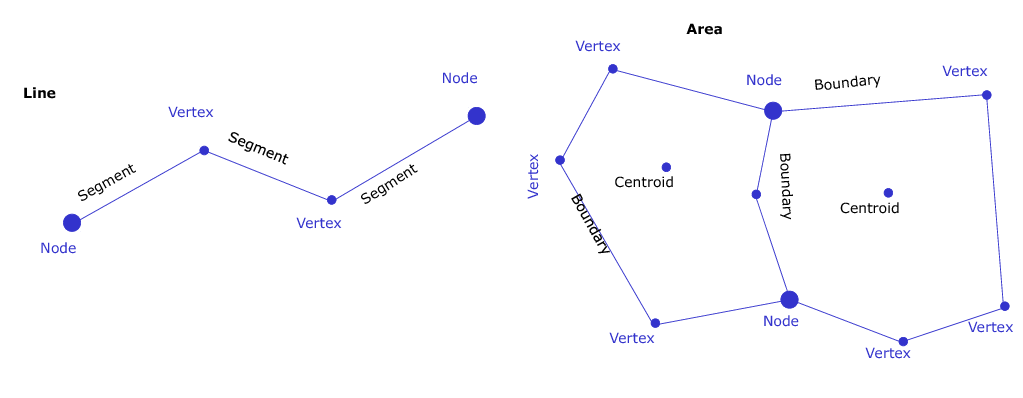
All vector data types in GRASS GIS
Modules
GRASS GIS is composed of more than 500 modules to perform any kind of GIS analysis. It is possible to analize raster, raster 3D, imagery and vector maps along with their alphanumerical attributes.The wealth of modules is organized by their first name in order to easily find the desired functionality. The graphical user interface offers a tree view as well as a search engine.
| Prefix | Function class | Type of command | Example |
|---|---|---|---|
| g.* | general | general data management | g.rename: renames map |
| d.* | display | graphical output | d.rast: display raster map, d.vect: display vector map |
| r.* | raster | raster processing | r.mapcalc: map algebra, r.univar: univariate statistics |
| v.* | vector | vector processing | v.clean: topological cleaning |
| i.* | imagery | imagery processing | i.pca: Principal Components Analysis on imagery group |
| r3.* | voxel | 3D raster processing | r3.stats: voxel statistics |
| db.* | database | database management | db.select: select value(s) from table |
| ps.* | postscript | map creation in PostScript format | ps.map: PostScript map creation |
| t.* | temporal | space-time datasets | t.rast.aggregate: raster time series aggregation |
It is also possible to install further modules, called Add-ons, from a centralized GRASS GIS Add-on repository at OSGeo or from Github using the command g.extension.
Region and Mask
The computational (or current) region is the actual setting of the region boundaries and the actual raster resolution.The raster input maps are automatically (on the fly) cropped/padded and rescaled to match the current region, the output maps have their bounds and resolution equal to those of the current computational region, while vector maps are always considered completely.
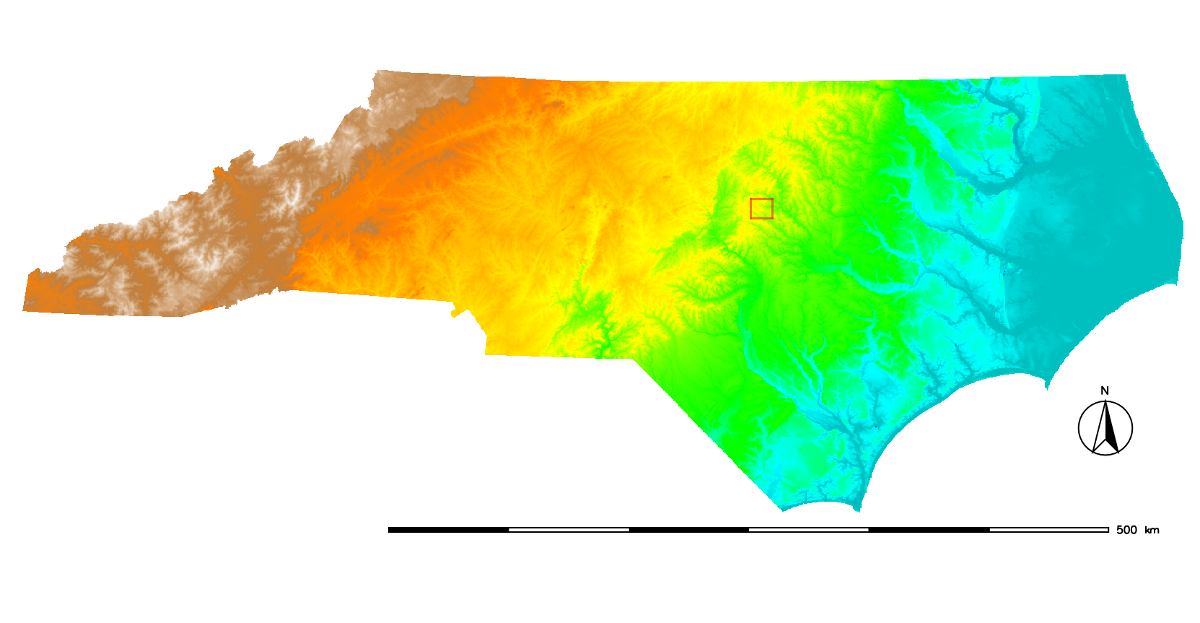
The red box shows the computational region currently set
MASK (this raster map name is reserved for this purpose).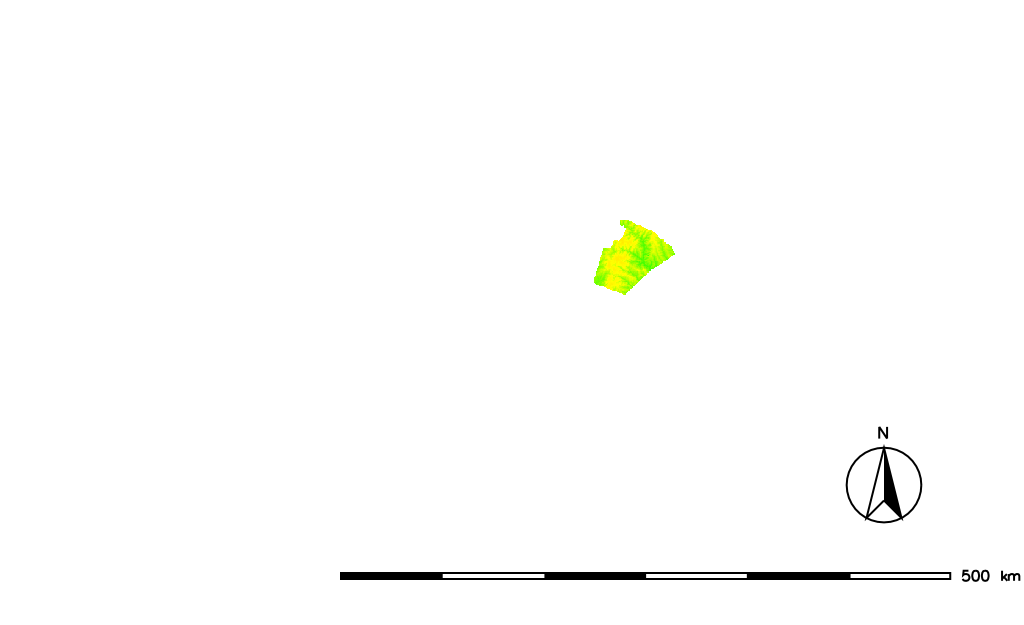
By setting the MASK (here based on a ZIP code) only the raster data inside the masked area are used for further analysis
Interfaces
GRASS GIS offers different interfaces for the interaction between user and software. Let's see them...Graphical User Interface
The GUI is the simpler way to approach GRASS GIS. The GRASS GIS GUI is composed of two elements, theLayer Manager where you can find all the GRASS GIS modules and manage your data and, the Map Display
where you can navigate, print and query your maps. The GUI also comes with a Python shell for rapid prototyping.
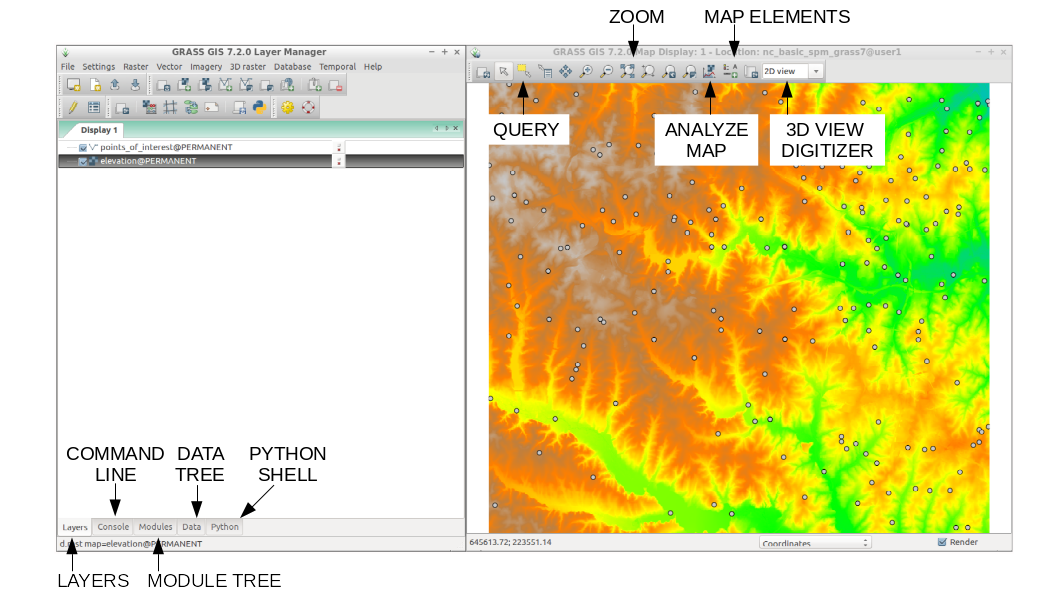
GRASS GIS Graphical User Interface (GUI)
QGIS
Otherwise you can also use GRASS GIS inside QGIS by either using theGRASS GIS plugin or through Processing.
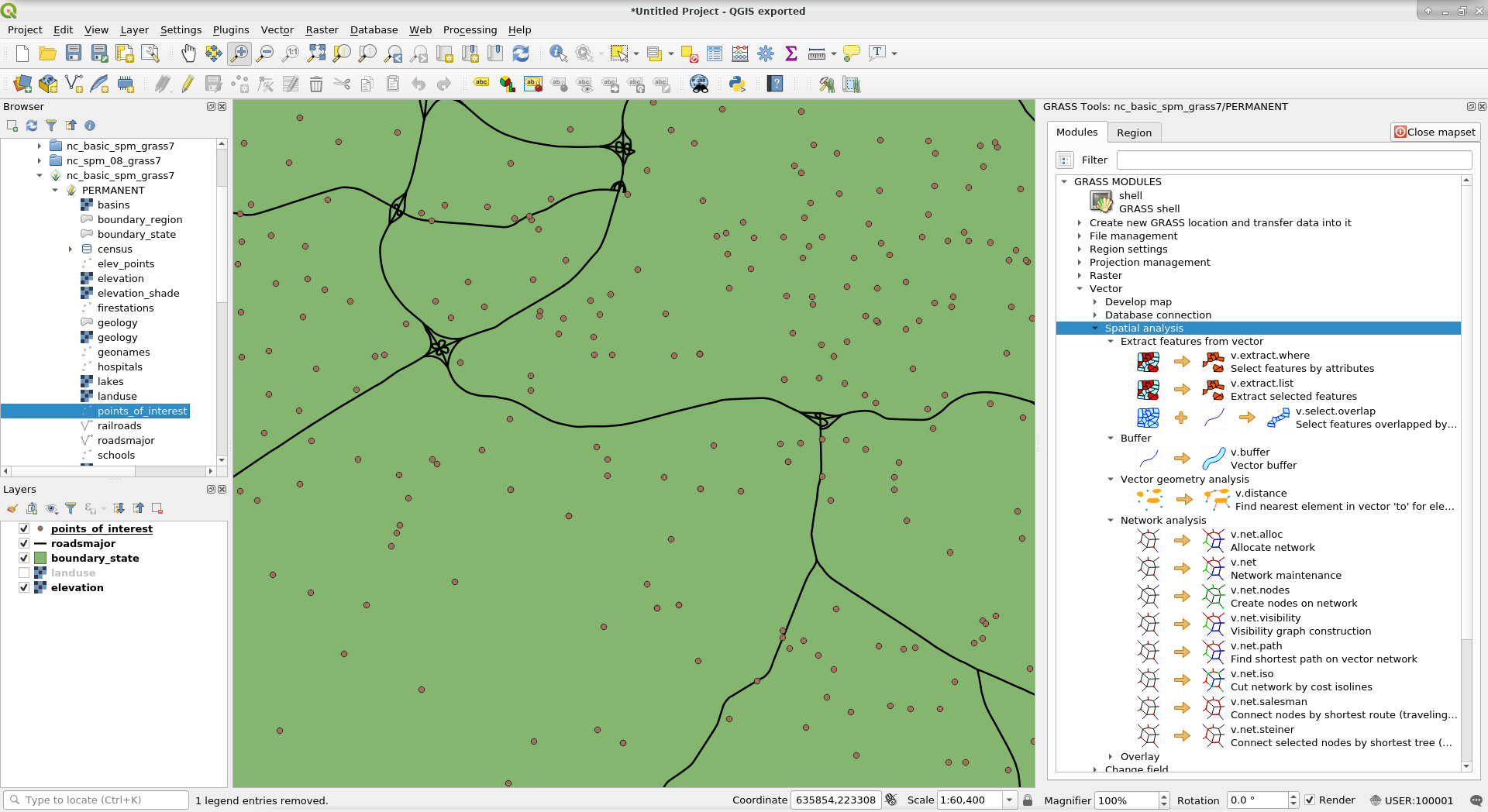
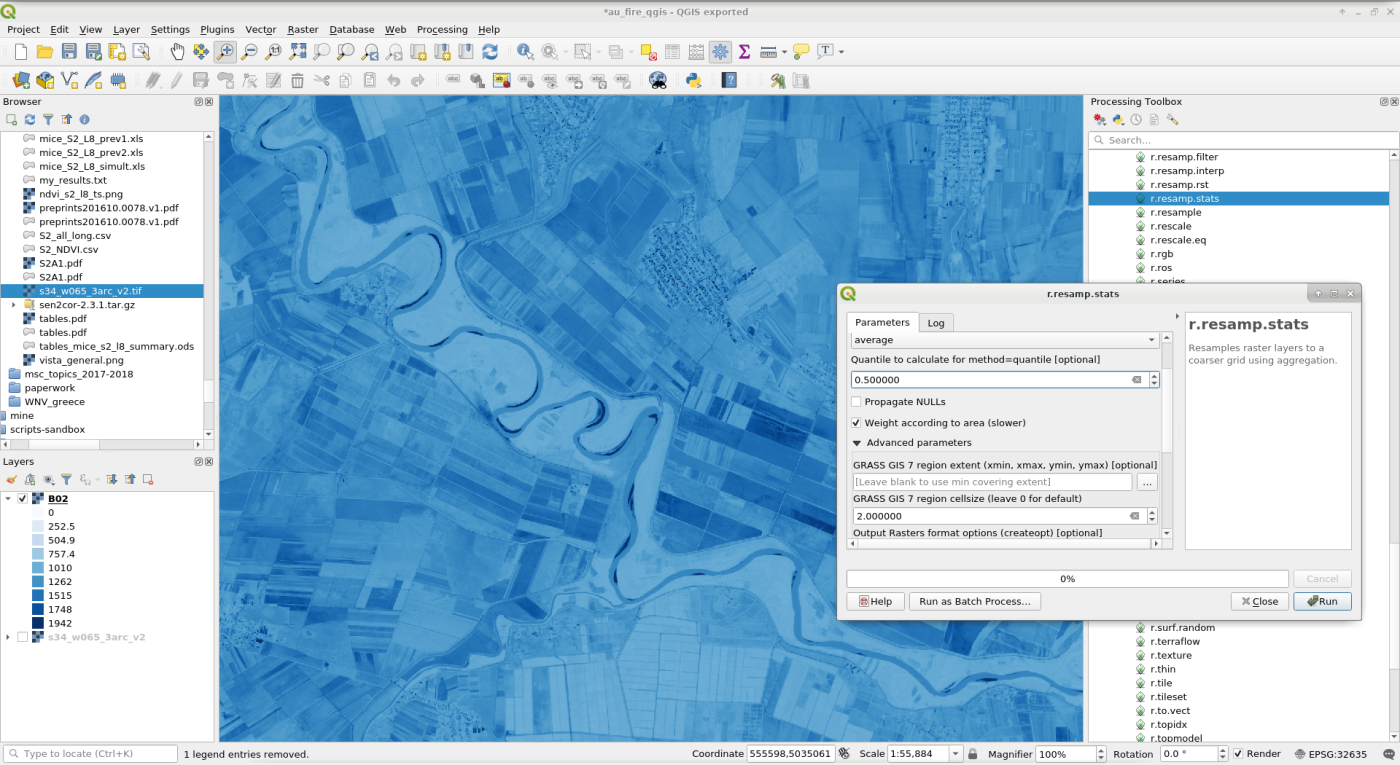
Using GRASS GIS within QGIS: GRASS GIS plugin (above), Processing toolbox (below).
Command line
The command line is the traditional and, probably, the most powerful way to use GRASS GIS, used daily by many GRASS GIS power users worldwide.Python
Python is a powerful and simple programming language, and you can use it to:- interface with the functionalities offered by GRASS GIS
- create your own workflows chaining several GRASS GIS modules
- create new add-ons by using GRASS GIS modules along with a wide number of Python libraries.
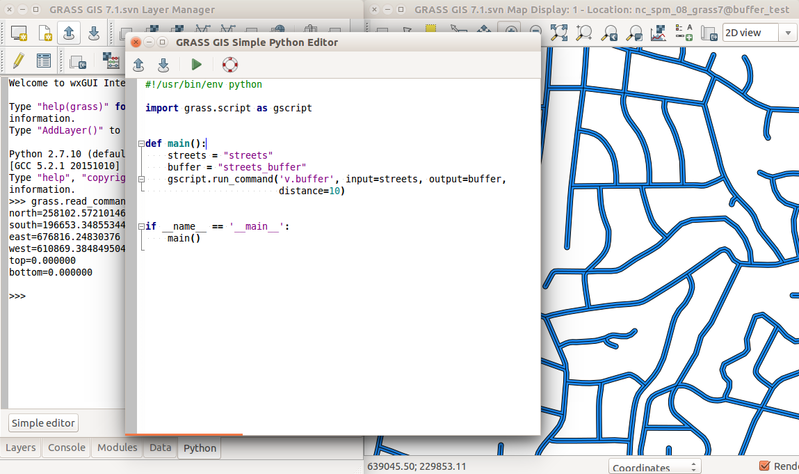
Simple Python editor in the GRASS GIS Graphical User Interface (GUI)
!/usr/bin/env python
# simple example for pyGRASS usage: raster processing via modules approach
from grass.pygrass.modules.shortcuts import general as g
from grass.pygrass.modules.shortcuts import raster as r
g.message("Filter elevation map by a threshold...")
# set computational region
input = 'elevation'
g.region(rast=input)
# hardcoded:
# r.mapcalc('elev_100m = if(elevation > 100, elevation, null())', overwrite = True)
# with variables
output = 'elev_100m'
thresh = 100.0
r.mapcalc("%s = if(%s > %d, %s, null())" % (output, input, thresh, input), overwrite = True)
r.colors(map=output, color="elevation")
R
R is a programming language and free software environment for statistical computing and graphics. You can use R in GRASS GIS in two different ways:- R within a GRASS GIS session
- GRASS GIS within an R session
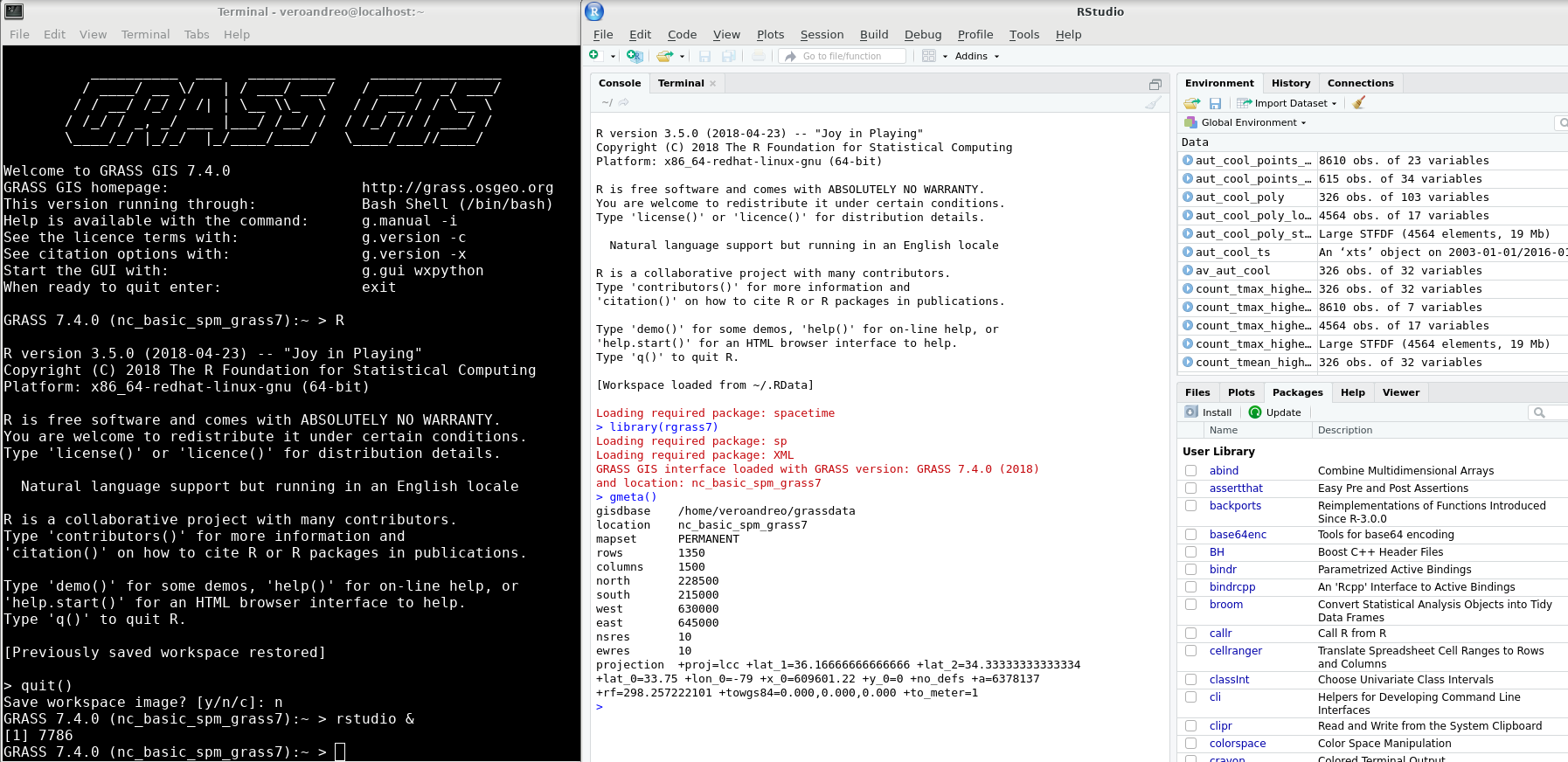
R within GRASS GIS console (left) and opening RStudio from GRASS GIS (right)
WPS - OGC Web Processing Service
It is possible to run GRASS GIS modules through the web using the Web Processing Service WPS is an OGC standard). The Free and Open Source software ZOO-Project and PyWPS allow the user to run GRASS GIS commands in a simple way.Temporal GRASS GIS introduction
GRASS GIS is the first Open Source GIS that incorporated capabilities to manage, analyze, process and visualize spatio-temporal data, as well as the temporal relationships among time series.
Importantly, the TGRASS concept is based on metadata and does not duplicate any datasets. It follows a snapshot approach, i.e.: add time stamps to existent maps, to integrate time dimension to the classical spatial GIS. In GRASS GIS words, all pixels in a raster map and all elements in a vector map, share the same time stamp. A collection of time stamped maps (snapshots) of the same variable are called space-time datasets (STDS) in TGRASS. Each map in a STDS might have a different spatial and temporal extent.
TGRASS uses an SQL database to store the temporal and spatial extension of STDS, as well as the topological relationships among maps and among STDS.
Terminology overview
Temporal database
A temporal database is a mapset-specific database in which all time stamped maps are registered, i.e.: their spatial and temporal extents, unique ID and map type metadata are stored. This allows users to perform complex SQL queries using the spatio-temporal extent and metadata information for map selection (See Temporal data analysis section).Space-time datasets
Space-time datasets are called differently according to the map type they are formed of:- Space time raster datasets (STRDS) collections of time stamped raster maps.
- Space time 3D raster datasets (STR3DS) collections of time stamped 3D raster maps.
- Space time vector datasets (STVDS) collections of time stamped vector maps.
Spatio-temporal modules
- t.*: General modules to handle STDS of all types
- t.rast.*: Modules that specifically deal with STRDS
- t.rast3d.*: Modules that specifically deal with STR3DS
- t.vect.*: Modules that specifically deal with STVDS
Absolute time vs relative time
Two different time definitions are used in TGRASS:
| Absolute time | Relative time |
|---|---|
| Gregorian calendar (ISO 8601 time format notation) | An integer and a time unit (year, month, day, hour, minute, seconds) |
| Example: 2013-10-15 13:00:00 | Example: 4 years, 90 days, 1 second |
Time intervals vs time instances
GRASS GIS supports both time intervals and time instances as map time stamps:
| Time intervals | Time instances |
|---|---|
| Defined by start time and end time: [start, end) | Defined by start time |
| Support for gaps, allow overlapping, might contain time instances, might be irregularly spaced in time | Punctual events |
Granularity
The granularity is the largest common divider granule of time intervals and gaps between intervals or instances from all time stamped maps that are collected in a STDS. It is represented as a number of seconds, minutes, hours, days, weeks, months or years.
Workflow overview
Now, how do we work with time series in GRASS GIS? Where do we start? Well, assuming we already have our maps in the GRASSDBASE, the first step is to set the connection to the temporal database. This is something we will need to do only once per mapset. After that, we create the STDSs, i.e.: the containers for the time series, and finally we assign time stamps to maps and register them in the STDS. After these basic steps, the following will depend on our specific objectives.- Set and connect the temporal database (mapset specific): t.connect
- Create the STDS (raster, vector or raster 3D): t.create
- Assign time stamps to maps and register them in the STDS: t.register
- Check basic info, integrity and validity of STDS: t.list, t.info, t.topology
- Edit, update, unregister, remove maps and/or STDS: t.rename, t.remove, t.support, t.unregister
- List maps, make selections, get univariate statistics: t.rast.list, t.vect.list, t.select, t.rast.extract, t.vect.extract, t.rast.univar, t.vect.univar
- Spatio-temporal processing (a bit raster biased):
- data aggregation: t.rast.series, t.rast.aggregate, t.rast.aggregate.ds
- data accumulation: t.rast.accumulate, t.rast.accumulate,
- gap-filling and smoothing: t.rast.gapfill, t.rast.neighbors
- spatio-temporal algebra: t.rast.mapcalc, t.rast.algebra, t.vect.algebra
- Visualization: g.gui.timeline, g.gui.tplot, g.gui.animation, g.gui.mapswipe
Hands-on to raster time series processing
Getting the MODIS satellite sensor data
MODIS is a payload scientific instrument on board the NASA Terra and the Aqua satellites with 36 spectral bands. Data are available for download upon user registration.
Create the SETTING file with the following content, e.g. in the directory $HOME/gisdata/:
your_NASA_user
your_NASA_password
The i.modis.download directly downloads from the MODIS data server into a local folder previously created (for example: lst_modis):
i.modis.download -g settings=$HOME/gisdata/SETTING product=lst_terra_monthly_5600 tiles=h11v05 startday="2015-01-01" endday="2016-12-31" folder=$HOME/lst_monthly
i.modis.import -w files=$HOME/lst_monthly/listfileMOD11B3.006.txt spectral="( 1 )" outfile=$HOME/lst_monthly/monthly_lst_to_register.txt
g.list type=raster pattern="MOD11B3*"
# Create folder to store .hdf files
mkdir $HOME/lst_monthly
# Download data: MOD11B3 2015-2016, tile h11v05
i.modis.download -g settings=$HOME/gisdata/SETTING \
product=lst_terra_monthly_5600 \
tiles=h11v05 startday="2015-01-01" endday="2016-12-31" \
folder=$HOME/lst_monthly
# Reproject from Sinusoidal to current NC projection and import maps into NC Location (Day and Night overpasses)
i.modis.import -w files=$HOME/lst_monthly/listfileMOD11B3.006.txt \
spectral="( 1 )" \
outfile=$HOME/lst_monthly/monthly_lst_to_register.txt
# Check list of imported maps
g.list type=raster pattern="MOD11B3*"
# Import the needed libraries
import grass.pygrass.modules as gmod
import os
from subprocess import PIPE
# Set variables
# home
HOME=os.path.expanduser('~')
# Create folder to store .hdf files
os.makedirs(os.path.join(HOME, lst_monthly))
# Download data: MOD11B3 2015-2016, tile h11v05
gmod.Module("i.modis.download", flags="g", settings=os.path.join(HOME, 'gisdata', 'SETTING'),
product="lst_terra_monthly_5600", tiles="h11v05", startday="2015-01-01",
endday="2016-12-31", folder=os.path.join(HOME, lst_monthly)
# Reproject and import maps into NC Location (Day and Night)
gmod.Module("i.modis.import", flags="w", files=os.path.join(HOME, 'lst_monthly', 'listfileMOD11B3.006.txt'),
spectral="( 1 )", outfile=os.path.join(HOME, 'lst_monthly', 'monthly_lst_to_register.txt'))
# Check list of imported maps
glist = gmod.Module("g.list", type="raster", pattern="MOD11B3*", stdout_=PIPE, stderr_=PIPE)
print(glist.outputs["stdout"].value)
We will now visualize one of the MODIS LST images and add different map decorations. We first change the color palette from grey to viridis and set the computational region to the state of North Carolina (using boundary_state vector map).
r.colors MOD11B3.A2015060.h11v05.single_LST_Day_6km color=viridis
g.region -p vector=boundary_state
# Change color palette from grey to viridis
r.colors MOD11B3.A2015060.h11v05.single_LST_Day_6km color=viridis
# Set region to NC (g.region -d)
g.region -p vector=boundary_state
# Optionally, to the full extent of one of the raster maps
g.region -p raster=MOD11B3.A2015060.h11v05.single_LST_Day_6km
# Change color palette from grey to viridis
gmod.Module("r.colors", map="MOD11B3.A2015060.h11v05.single_LST_Day_6km", color="viridis")
# Set region to NC
gmod.Module("g.region", flags="p", vector="boundary_state")
# Optionally, to the full extent of one of the raster maps
gmod.Module("g.region", flags="p", raster="MOD11B3.A2015060.h11v05.single_LST_Day_6km")
We can open a monitor and run the commands from the command line or do everything in the main GUI and copy the commands for future reference or replication. Note the Copy button in each command GUI.
# Open a monitor
d.mon wx0
# Display raster map
d.rast map=MOD11B3.A2015060.h11v05.single_LST_Day_6km
# Display vector map
d.vect map=boundary_state type=boundary
# Add raster legend
d.legend -t -s -b raster=MOD11B3.A2015060.h11v05.single_LST_Day_6km \
title=LST title_fontsize=20 font=sans fontsize=18
# Add scale bar
d.barscale length=200 units=kilometers segment=4 fontsize=14
# Add North arrow
d.northarrow style=1b text_color=black
# Add text
d.text -b text="LST Day from MOD11B3.006 - North Carolina - March, 2015" \
color=black bgcolor=229:229:229 align=cc font=sans size=8
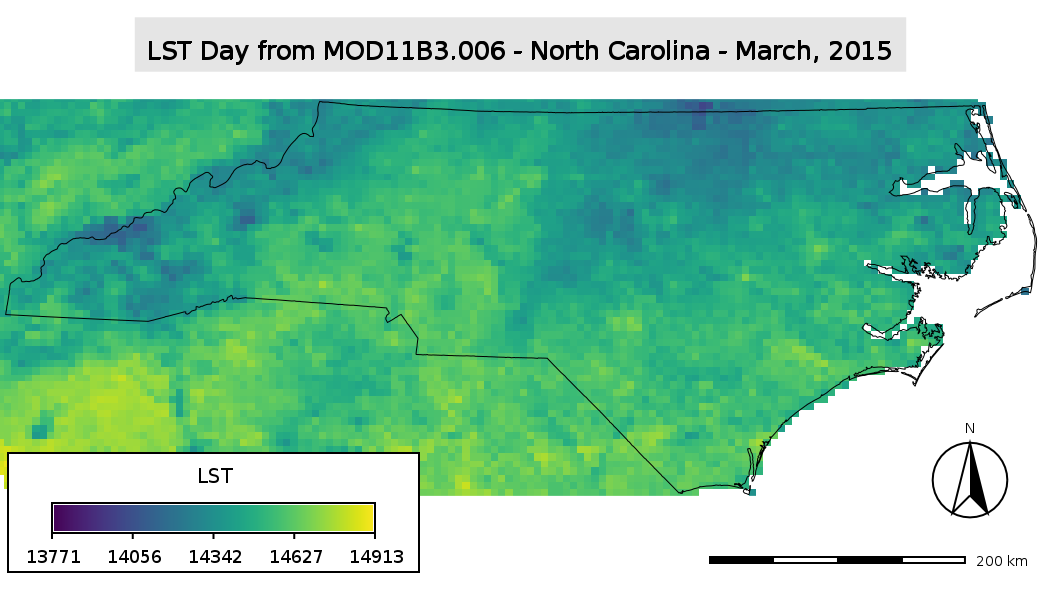
MODIS LST map with decorations (scaled Kelvin pixel values)
Create temporal raster dataset (STRDS)
To create a space-time raster data set (STRDS) implies to create an SQLite table in the temporal database, i.e.: a container table, that will hold our raster time series and will allow us to easily handle huge amounts of maps by only using the STRDS as input in temporal commands.
If this is the first time you use the temporal framework, you need to create and set the connection to the temporal database by means of t.connect. As the temporal database is mapset specific, you'll need to repeat this step in each mapset in which you'll have STDSs.
Once the connection is set, you can create the empty STRDS, i.e.: the empty table, in which you'll put (aka: register) all your time series maps afterwards. For the creation of any STDS, we need to specify which type of maps (raster, raster3d or vector) the STDS will contain and which type of time (absolute or relative) the maps represent.
t.connect -d
t.create type=strds temporaltype=absolute output=LST_Day_monthly title="Monthly LST Day 5.6 km" description="Monthly LST Day 5.6 km MOD11B3.006, 2015-2016"
t.list type=strds
t.info input=LST_Day_monthly
# Create the temporal db connection
t.connect -d
# Create the STRDS
t.create type=strds temporaltype=absolute output=LST_Day_monthly \
title="Monthly LST Day 5.6 km" \
description="Monthly LST Day 5.6 km MOD11B3.006, 2015-2016"
# Check if the STRDS is created
t.list type=strds
# Get info about the STRDS
t.info input=LST_Day_monthly
import grass.temporal as tgis
# Create the temporal db connection using temporal library
gmod.Module("t.connect", flags="d")
# Initialize the temporal library
tgis.init()
# Create the new STRDS
tgis.open_new_stds("LST_Day_monthly", "strds", "absolute", "Monthly LST Day 5.6 km",
"Monthly LST Day 5.6 km MOD11B3.006, 2015-2016", "mean", None, False)
# Connect to the SQL database interface
dbif = tgis.SQLDatabaseInterfaceConnection()
dbif.connect()
# Get and print the list of temporal dataset
stds_list = tgis.get_dataset_list("strds", "absolute", dbif=dbif)
print(stds_list)
# Get and print info about the created strds
dataset = tgis.dataset_factory("strds", "LST_Day_monthly@modis_lst")
dataset.select(dbif)
dataset.print_info()
Once the STRDS is created, we assign time stamps to maps and add them to the STRDS, i.e.: we register maps in the STRDS. To register maps in a STDS, we need to pass the empty STDS as input and the list of maps to be registered. There are different ways to register maps in STDS. For more options, you can check the t.register manual and the related wiki page.
t.register input=LST_Day_monthly file=$HOME/lst_monthly/monthly_lst_to_register.txt
# Register maps in STRDS
t.register input=LST_Day_monthly \
file=$HOME/lst_monthly/monthly_lst_to_tregister.txt
# Register maps using the Python function
gmod.Module("t.register", input="LST_Day_monthly@modis_lst",
file=os.path.join(HOME, 'lst_monthly', 'monthly_lst_to_register.txt'))
Alternatively, we could have registered our raster maps using the maps, start and increment options along with the i flag for interval creation. The command in that case would look as follows:
t.register -i input=LST_Day_monthly \
maps=`g.list type=raster pattern=MOD11B3*LST_Day* separator=comma` \
start="2015-01-01" increment="1 months"
Let's check now the basic info again to see how it looks like and list the raster maps in our LST_Day_monthly STRDS:
t.info LST_Day_monthly
t.rast.list LST_Day_monthly
# Check info
t.info LST_Day_monthly
# Check the list of maps in the STRDS
t.rast.list LST_Day_monthly
# Get info using the modules
gmod.Module("t.info", input="LST_Day_monthly")
# Get list of raster using the Python library
tgis.list_maps_of_stds("strds", "LST_Day_monthly@modis_lst", "name,mapset,start_time,end_time",
"start_time", "", "|", "cols", True, "no")
Let's see our STRDS graphically. We will use the g.gui.timeline tool.
g.gui.timeline inputs=LST_Day_monthly
The tool g.gui.timeline is under the Temporal menu, in GUI Tools section. It can also be called from the terminal with: g.gui.timeline --ui.
To plot a STDS:
1. Choose a STRDS from the dropdown list.
2. Click the Draw button.
3. Change the margin settings as desired.
Alternatively, you can also get a 3D representation of the STDS (i.e.: x, y and time) by checking the 3D box.
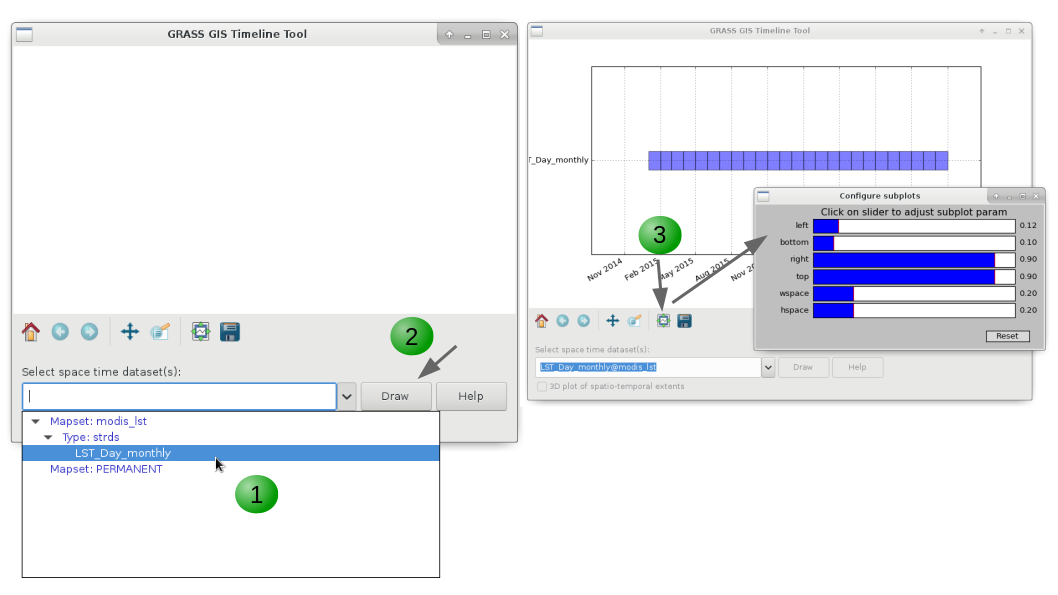
Workflow for plotting STRDS with g.gui.timeline
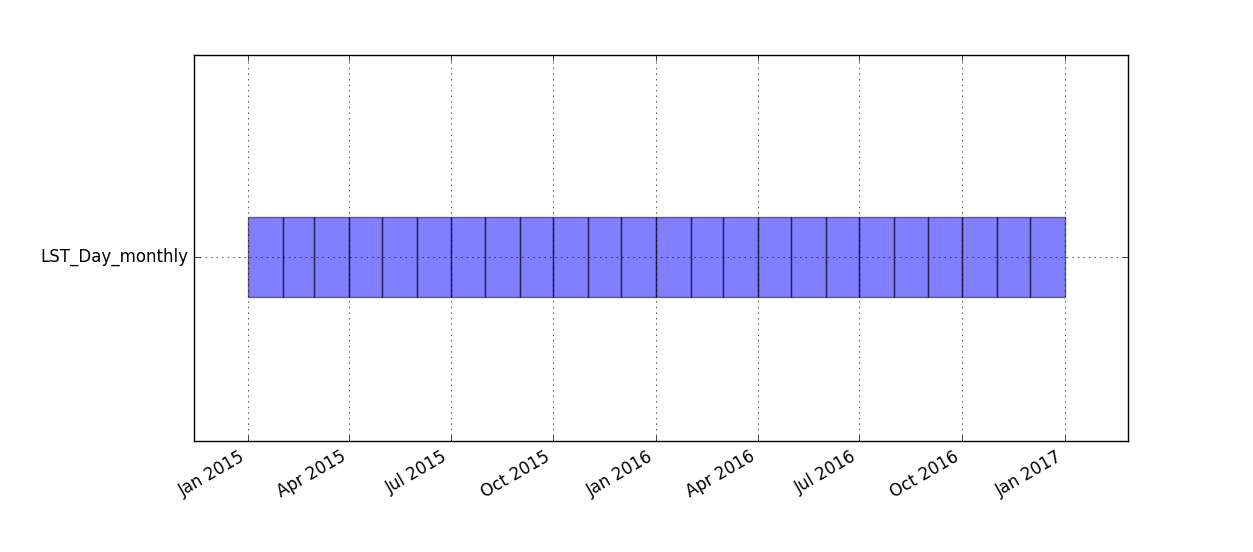
Timeline plot for LST_Day_monthly time series
Temporal data analysis
One basic but very important function when handling hundreds or thousands of maps is the listing function, i.e.: we usually need to list maps that meet a certain condition. For example, we need maps which start month is June, maps with minimum values lower than 100, and so on. The GRASS GIS Temporal framework has different commands for that task: t.list for listing STDS and maps registered in the temporal database, t.rast.list for maps in raster time series and, t.vect.list for maps in vector time series. All these commands allow us to list STDSs and/or maps according to different criteria. Let's see some examples with our LST_Day_monthly STRDS:
# Maps with minimum value lower than or equal to 14000
t.rast.list input=LST_Day_monthly order=min columns=name,start_time,min where="min <= '14000'"
name|start_time|min
MOD11B3.A2015032.h11v05.single_LST_Day_6km|2015-02-01 00:00:00|12950.0
MOD11B3.A2016032.h11v05.single_LST_Day_6km|2016-02-01 00:00:00|12964.0
MOD11B3.A2015001.h11v05.single_LST_Day_6km|2015-01-01 00:00:00|13022.0
# Maps with maximum value higher than 14000
t.rast.list input=LST_Day_monthly order=max columns=name,start_time,max where="max > '14000'"
name|start_time|max
MOD11B3.A2016001.h11v05.single_LST_Day_6km|2016-01-01 00:00:00|14360.0
MOD11B3.A2015001.h11v05.single_LST_Day_6km|2015-01-01 00:00:00|14396.0
MOD11B3.A2015032.h11v05.single_LST_Day_6km|2015-02-01 00:00:00|14522.0
# Maps between two given dates
t.rast.list input=LST_Day_monthly columns=name,start_time where="start_time >= '2015-05' and start_time <= '2015-08-01 00:00:00'"
name|start_time
MOD11B3.A2015121.h11v05.single_LST_Day_6km|2015-05-01 00:00:00
MOD11B3.A2015152.h11v05.single_LST_Day_6km|2015-06-01 00:00:00
MOD11B3.A2015182.h11v05.single_LST_Day_6km|2015-07-01 00:00:00
MOD11B3.A2015213.h11v05.single_LST_Day_6km|2015-08-01 00:00:00
# Maps from January
t.rast.list input=LST_Day_monthly columns=name,start_time where="strftime('%m', start_time)='01'"
name|start_time
MOD11B3.A2015001.h11v05.single_LST_Day_6km|2015-01-01 00:00:00
MOD11B3.A2016001.h11v05.single_LST_Day_6km|2016-01-01 00:00:00
# Maps from June, 1st
t.rast.list input=LST_Day_monthly columns=name,start_time where="strftime('%m-%d', start_time)='06-01'"
name|start_time
MOD11B3.A2015152.h11v05.single_LST_Day_6km|2015-06-01 00:00:00
MOD11B3.A2016153.h11v05.single_LST_Day_6km|2016-06-01 00:00:00
To explore a bit more our time series, we will obtain univariate statistics for the maps in the STRDS. There's a dedicated module for that: t.rast.univar. There's also the possibility to obtain extended statistics such as first quartile, median value, third quartile and percentile 90 by setting the e flag. Since we start to run analysis it is better to set the computational region to match with the rasters. Let's see:
g.region raster=MOD11B3.A2015001.h11v05.single_LST_Day_6km@modis_lst -p
t.rast.univar input=LST_Day_monthly
id|start|end|mean|min|max|mean_of_abs|stddev|variance|coeff_var|sum|null_cells|cells
MOD11B3.A2015001.h11v05.single_LST_Day_6km@modis_lst|2015-01-01 00:00:00|2015-02-01 00:00:00|13920.4954589617|13022|14396|13920.4954589617|204.922713367766|41993.3184540075|1.47209353267553|329539889|51297|74970
MOD11B3.A2015032.h11v05.single_LST_Day_6km@modis_lst|2015-02-01 00:00:00|2015-03-01 00:00:00|13850.5832593907|12950|14522|13850.5832593907|269.511406162359|72636.3980516119|1.94584878567933|327801754|51303|74970
MOD11B3.A2015060.h11v05.single_LST_Day_6km@modis_lst|2015-03-01 00:00:00|2015-04-01 00:00:00|14387.8026526992|13771|14913|14387.8026526992|202.223782537869|40894.4582239235|1.40552235403321|340616840|51296|74970
MOD11B3.A2015091.h11v05.single_LST_Day_6km@modis_lst|2015-04-01 00:00:00|2015-05-01 00:00:00|14724.7657768016|14160|15212|14724.7657768016|127.152673621415|16167.8024090742|0.863529346060911|348594105|51296|74970
t.rast.univar -e input=LST_Day_monthly
id|start|end|mean|min|max|mean_of_abs|stddev|variance|coeff_var|sum|null_cells|cells|first_quartile|median|third_quartile|percentile_90
MOD11B3.A2015001.h11v05.single_LST_Day_6km@modis_lst|2015-01-01 00:00:00|2015-02-01 00:00:00|14063.8152535675|13625|14289|14063.8152535675|104.326695294161|10884.0593510007|0.741809341300163|12313517190|135054|1010600|14001|14076|14148|14183
MOD11B3.A2015032.h11v05.single_LST_Day_6km@modis_lst|2015-02-01 00:00:00|2015-03-01 00:00:00|14040.1111911881|13532|14349|14040.1111911881|128.052778812874|16397.5141616988|0.912049606083195|12292763193|135054|1010600|13944|14059|14140|14190
MOD11B3.A2015060.h11v05.single_LST_Day_6km@modis_lst|2015-03-01 00:00:00|2015-04-01 00:00:00|14509.6026765013|14012|14884|14509.6026765013|128.58386990137|16533.8115988123|0.886198421612294|12703824585|135054|1010600|14417|14523|14599|14664
MOD11B3.A2015091.h11v05.single_LST_Day_6km@modis_lst|2015-04-01 00:00:00|2015-05-01 00:00:00|14781.4240850852|14160|15128|14781.4240850852|116.594077815457|13594.1789816369|0.788787853892261|12941816732|135054|1010600|14724|14802|14859|14907
t.rast.univar input=LST_Day_monthly separator=comma output=stats_LST_Day_monthly.csv
# Set the computational region match to one of the raster
g.region raster=MOD11B3.A2015001.h11v05.single_LST_Day_6km@modis_lst -p
# Print univariate stats for maps within STRDS
t.rast.univar input=LST_Day_monthly
# Get extended statistics
t.rast.univar -e input=LST_Day_monthly
# Write the univariate stats output to a csv file
t.rast.univar input=LST_Day_monthly separator=comma output=stats_LST_Day_monthly.csv
# Set the computational region match to one of the raster
gmod.Module("g.region", flags="p", raster="MOD11B3.A2015001.h11v05.single_LST_Day_6km@modis_lst")
# Print univarite statistics with the Python library
tgis.print_gridded_dataset_univar_statistics("strds", "LST_Day_monthly@modis_lst", None, None, False, False, ",", False)
# Print extended univarite statistics with the Python library
tgis.print_gridded_dataset_univar_statistics("strds", "LST_Day_monthly@modis_lst", None, None, True, False, ",", False)
However, those statistics are a bit difficult to directly interpret since values are in °K*50. Let's re-scale the data to °C and run the previous command again. Then, we will set a proper color palette for the STRDS and display one map. The latter, you already know how to do.
To transform all the maps in our LST_Day_monthly time series into °C we will use the t.rast.algebra module. This module allows us to perform a very wide variety of operations in the temporal and spatial domains, as well as much of the more "classical" operations already available in r.mapcalc.
t.rast.algebra basename=LST_Day_monthly_celsius expression="LST_Day_monthly_celsius = LST_Day_monthly * 0.02 - 273.15"
t.rast.univar input=LST_Day_monthly_celsius
t.rast.colors input=LST_Day_monthly_celsius color=celsius
# Re-scale data into degrees Celsius
t.rast.algebra basename=LST_Day_monthly_celsius \
expression="LST_Day_monthly_celsius = LST_Day_monthly * 0.02 - 273.15"
# Print univariate stats
t.rast.univar input=LST_Day_monthly_celsius
# Set a dedicated color pallete
t.rast.colors input=LST_Day_monthly_celsius color=celsius
# Re-scale data into degrees Celsius
gmod.Module("t.rast.algebra", basename="LST_Day_monthly_celsius",
expression="LST_Day_monthly_celsius = LST_Day_monthly * 0.02 - 273.15")
# Print univarite statistics with the Python library
tgis.print_gridded_dataset_univar_statistics("strds", "LST_Day_monthly_celsius@modis_lst", None, None, False, False, ",", False)
gmod.Module("t.rast.colors", input="LST_Day_monthly_celsius", color="celsius")
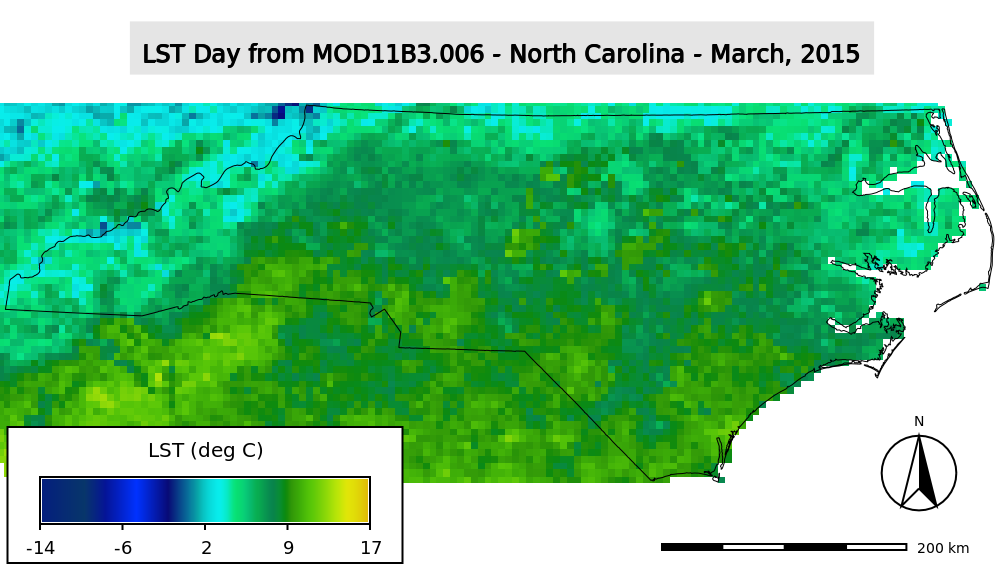
MODIS LST re-scaled to degrees Celsius
Alternatively, we could have used t.rast.mapcalc to perform the previous transformation. The command would look like this:
t.rast.mapcalc input=LST_Day_monthly output=LST_Day_monthly_celsius basename=LST_Day_monthly_celsius expression="LST_Day_monthly * 0.02 - 273.15"
Temporal aggregations
There are basically two dedicated modules to perform temporal aggregations in GRASS GIS. The first one that we will use is t.rast.series. This module is a wrapper for r.series and allows us to aggregate our STRDS or parts of it with different methods. We will use it now to obtain the absolute maximum LST in the past two years.
t.rast.series input=LST_Day_monthly_celsius output=LST_Day_max method=maximum
r.colors map=LST_Day_max color=celsius
d.mon wx0
d.rast LST_Day_max
d.vect map=boundary_state type=boundary
d.legend -t -s -b raster=LST_Day_max title=LST title_fontsize=20 font=sans fontsize=18
d.barscale length=200 units=kilometers segment=4 fontsize=14
d.northarrow style=1b text_color=black
d.text -b text="Maximum LST in the period 2015-2016 - North Carolina" color=black bgcolor=229:229:229 align=cc font=sans size=8
# Get maximum LST in the STRDS
t.rast.series input=LST_Day_monthly_celsius \
output=LST_Day_max method=maximum
# Change color pallete to celsius
r.colors map=LST_Day_max color=celsius
# Display the new map
d.mon wx0
d.rast LST_Day_max
d.vect map=boundary_state type=boundary
d.legend -t -s -b raster=LST_Day_max \
title=LST title_fontsize=20 font=sans fontsize=18
d.barscale length=200 units=kilometers segment=4 fontsize=14
d.northarrow style=1b text_color=black
d.text -b text="Maximum LST in the period 2015-2016 - North Carolina" \
color=black bgcolor=229:229:229 align=cc font=sans size=8
# Get maximum LST in the STRDS
gmod.Module("t.rast.series", input="LST_Day_monthly_celsius",
output="LST_Day_max", method="maximum")
# Change color pallete to celsius
gmod.Module("r.colors", map="LST_Day_max", color="celsius")
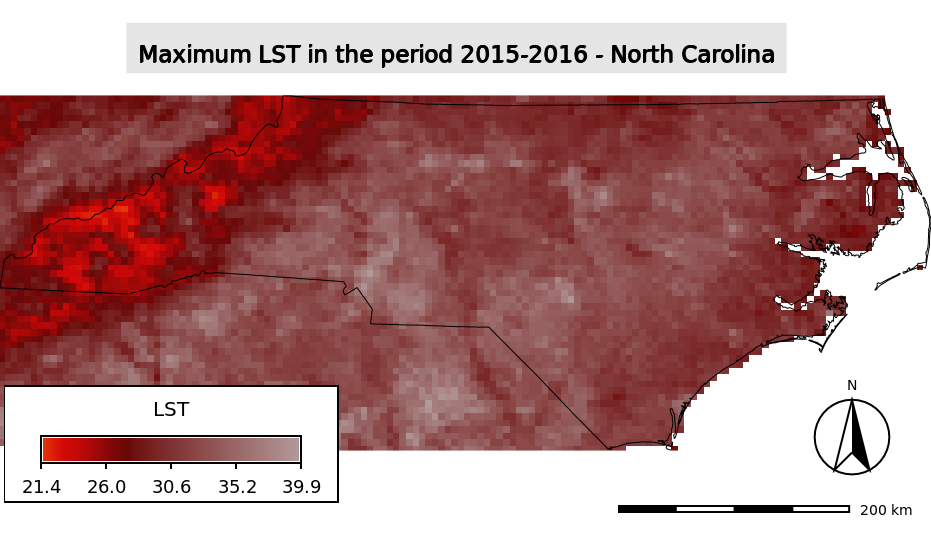
Maximum LST in the period 2015-2016 in North Carolina
Now, by means of the spatio-temporal map calculator, we will get the month in which the absolute maximum LST occurred. For that, we will first compare our LST_Day_monthly_celsius STRDS with the map of absolute maximum LST LST_Day_max that we just obtained before. If they coincide, we keep the month for that pixel, otherwise it will be NULL. Then, we aggregate the resulting month_max_lst STRDS with t.rast.series method=maximum and we get the map with the pixelwise month in which the absolute maximum LST has occurred in the past two years. Finally, we remove the intermediate STRDS, since we are only interested in the aggregated map.
t.rast.mapcalc -n inputs=LST_Day_monthly_celsius output=month_max_lst expression="if(LST_Day_monthly_celsius == LST_Day_max, start_month(), null())" basename=month_max_lst
t.rast.series input=month_max_lst method=maximum output=max_lst_date
t.remove -rf inputs=month_max_lst
# New strds with month of overall maximum
t.rast.mapcalc -n inputs=LST_Day_monthly_celsius output=month_max_lst \
expression="if(LST_Day_monthly_celsius == LST_Day_max, start_month(), null())" \
basename=month_max_lst
# Get basic info
t.info month_max_lst
# Map with month of overall maximum
t.rast.series input=month_max_lst method=maximum output=max_lst_date
# Remove month_max_lst strds (we were only interested in the resulting aggregated map)
t.remove -rf inputs=month_max_lst
# New strds with month of overall maximum
gmod.Module("t.rast.mapcalc", flags="n", inputs="LST_Day_monthly_celsius", output="month_max_lst",
expression="if(LST_Day_monthly_celsius == LST_Day_max, start_month(), null())",
basename="month_max_lst")
# Map with month of overall maximum
gmod.Module("t.rast.series", input="month_max_lst", method="maximum", output="max_lst_date")
# Remove month_max_lst strds
gmod.Module("t.remove", flags="rf", inputs="month_max_lst")
Note that the flags "-rf" force (immediate) removal of both the STRDS (i.e.: the container table) and the maps registered in it.
Finally, we display the resulting map:
d.mon wx0
d.rast max_lst_date
d.vect map=boundary_state type=boundary
d.legend -t -s -b raster=max_lst_date title=LST title_fontsize=20 font=sans fontsize=18
d.barscale length=200 units=kilometers segment=4 fontsize=14
d.northarrow style=1b text_color=black
d.text -b text="Month of maximum LST 2015-2016" color=black bgcolor=229:229:229 align=cc font=sans size=8
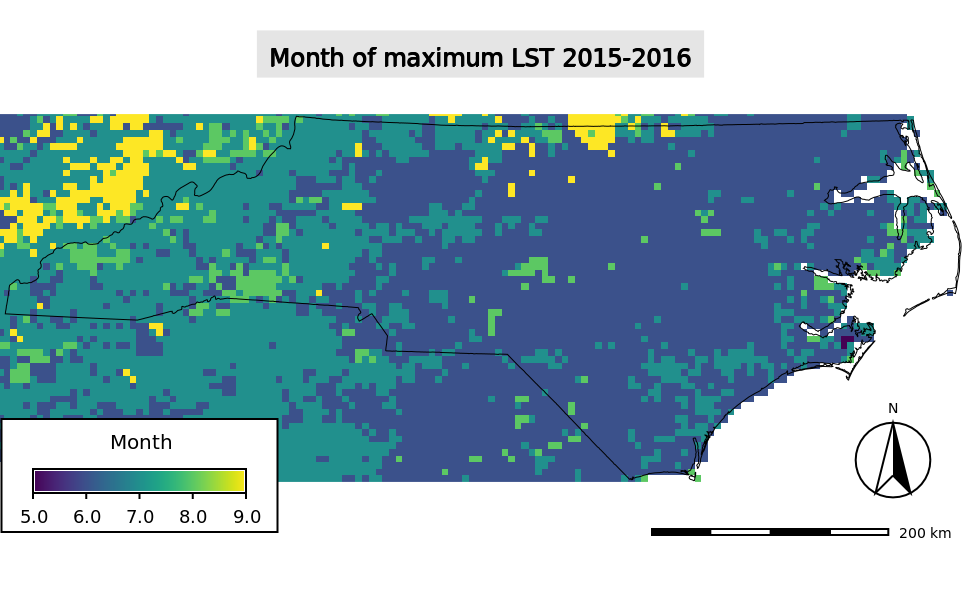
Month of maximum LST for the period 2015-2016
The other module that allows us to perform temporal aggregations is t.rast.aggregate. With this module we are able to aggregate raster maps in our STRDS with different granularities. Note that this module also has the option where that allows us to set specific dates for the aggregation. We will use this module to get 3-month and 6-month average LST.
t.rast.aggregate input=LST_Day_monthly_celsius output=LST_Day_mean_3month basename=LST_Day_mean_3month suffix=gran method=average granularity="3 months"
t.info LST_Day_mean_3month
t.rast.list LST_Day_mean_3month
t.rast.aggregate input=LST_Day_monthly_celsius output=LST_Day_mean_6month basename=LST_Day_mean_6month suffix=gran method=average granularity="6 months"
t.info LST_Day_mean_6month
t.rast.list LST_Day_mean_6month
# 3-month mean LST
t.rast.aggregate input=LST_Day_monthly_celsius \
output=LST_Day_mean_3month \
basename=LST_Day_mean_3month suffix=gran \
method=average granularity="3 months"
# Print info
t.info LST_Day_mean_3month
# Print list of maps
t.rast.list LST_Day_mean_3month
# 6-month mean LST
t.rast.aggregate input=LST_Day_monthly_celsius \
output=LST_Day_mean_6month \
basename=LST_Day_mean_6month suffix=gran \
method=average granularity="6 months"
# Print info
t.info LST_Day_mean_6month
# Print list of maps
t.rast.list LST_Day_mean_6month
# 3-month mean LST
gmod.Module("t.rast.aggregate", input="LST_Day_monthly_celsius", output="LST_Day_mean_3month",
basename="LST_Day_mean_3month", suffix="gran", method="average", granularity="3 months")
# Get and print info about LST_Day_mean_3month strds in shell output format
dataset = tgis.dataset_factory("strds", "LST_Day_mean_3month@modis_lst")
dataset.select(dbif)
dataset.print_shell_info()
# Print list of maps
tgis.list_maps_of_stds("strds", "LST_Day_mean_3month@modis_lst", "name,mapset,start_time,end_time",
"start_time", "", "|", "cols", True, "no")
# 6-month mean LST
gmod.Module("t.rast.aggregate", input="LST_Day_monthly_celsius", output="LST_Day_mean_6month",
basename="LST_Day_mean_6month", suffix="gran", method="average", granularity="6 months")
# Get and print info about LST_Day_mean_6month strds
gmod.Module("t.info", input="LST_Day_mean_6month")
# Print list of maps
gmod.Module("t.rast.list", input="LST_Day_mean_6month")
Now, we will extract the 3-month average LST for points in a vector map. We will use the vector map points_of_interest that is already available in the PERMANENT mapset of NC Location. There are different commands that allow us to perform this task, but we'll use t.rast.what. This module samples a STRDS at specific vector point coordinates and writes the output to stdout using different layouts. You might explore the options available in the parameter layout to see the different ways to write the output.
t.rast.what -n points=points_of_interest strds=LST_Day_mean_6month null_value=NA separator=comma
x,y,start,end,value
646341.7386813260,218873.7305680360,2015-01-01 00:00:00,2015-07-01 00:00:00,19.3766666666667
646341.7386813260,218873.7305680360,2015-07-01 00:00:00,2016-01-01 00:00:00,23.4166666666667
646341.7386813260,218873.7305680360,2016-01-01 00:00:00,2016-07-01 00:00:00,20.5866666666667
646341.7386813260,218873.7305680360,2016-07-01 00:00:00,2017-01-01 00:00:00,24.2266666666667
...
641015.3265248850,254295.3989810100,2015-01-01 00:00:00,2015-07-01 00:00:00,15.5833333333334
641015.3265248850,254295.3989810100,2015-07-01 00:00:00,2016-01-01 00:00:00,20.74
641015.3265248850,254295.3989810100,2016-01-01 00:00:00,2016-07-01 00:00:00,16.92
641015.3265248850,254295.3989810100,2016-07-01 00:00:00,2017-01-01 00:00:00,20.76
t.rast.what -n points=points_of_interest strds=LST_Day_mean_6month output=trastwhat_output.csv null_value=NA separator=comma
# Print STRDS values for the points with header (-n flag)
t.rast.what -n points=points_of_interest strds=LST_Day_mean_6month null_value=NA separator=comma
# Write STRDS values for the points to a csv file
t.rast.what -n points=points_of_interest strds=LST_Day_mean_6month output=trastwhat_output.csv \
null_value=NA separator=comma
# Print STRDS values for the points with header (-n flag)
gmod.Module("t.rast.what", flags="n", points="points_of_interest",
strds="LST_Day_mean_6month", null_value="NA", separator="comma")
# Write STRDS values for the points to a csv file
gmod.Module("t.rast.what", flags="n", points="points_of_interest",
strds="LST_Day_mean_6month", output="trastwhat_output.csv",
null_value="NA", separator="comma")
Even though the most common is to extract raster data to points, we may also be interested in getting spatially aggregated time series data for polygons. GRASS GIS has an add-on for this: v.strds.stats. It calculates zonal statistics of each raster in a STRDS and writes the output to the attribute table of a new polygon vector map. We first need to install the add-on through g.extension. Then, we will extract the average, minimum and maximum monthly LST for the different geologic types in NC.
g.extension v.strds.stats
v.strds.stats input=geology strds=LST_Day_monthly_celsius output=geology_aggr_lst method=average,minimum,maximum
v.db.select map=geology_aggr_lst file=ts_polygons.csv
# Install v.strds.stats add-on
g.extension v.strds.stats
# Extract mean, max and min LST for municipalities
v.strds.stats input=geology strds=LST_Day_monthly_celsius \
output=geology_aggr_lst method=average,minimum,maximum
# Save the attribute table of the new vector into a csv file
v.db.select map=geology_aggr_lst file=ts_polygons.csv
# Install v.strds.stats add-on
gmod.Module("g.extension", extension="v.strds.stats")
# Extract mean, max and min LST for municipalities
gmod.Module("v.strds.stats", input="geology", output="geology_aggr_lst",
strds="LST_Day_monthly_celsius", method="average,minimum,maximum")
# Save the attribute table of the new vector into a csv file
gmod.Module("v.db.select", map="geology_aggr_lst", file="ts_polygons.csv")
Temporal data visualization
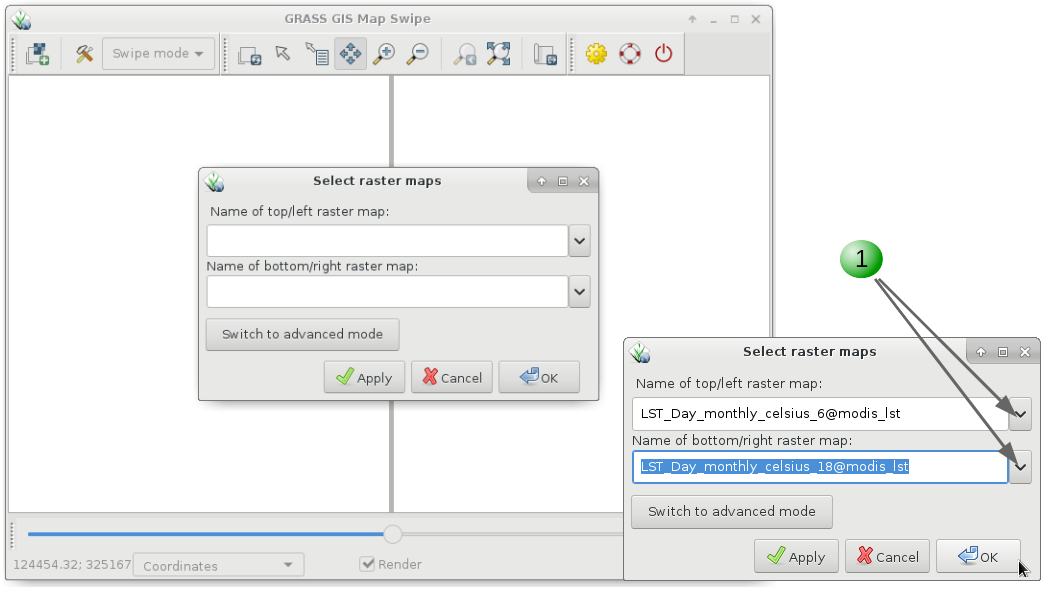
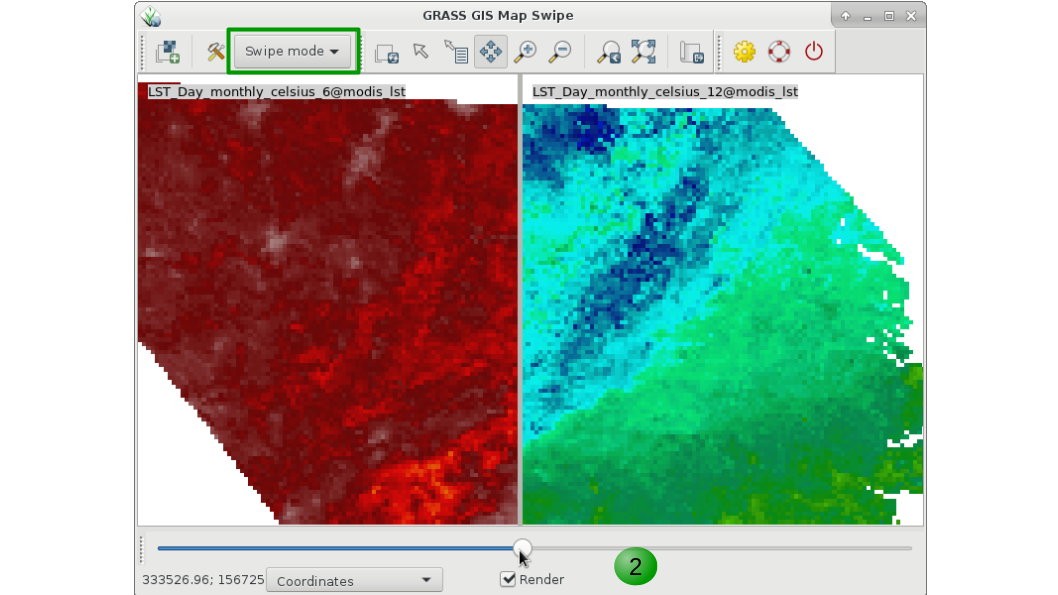
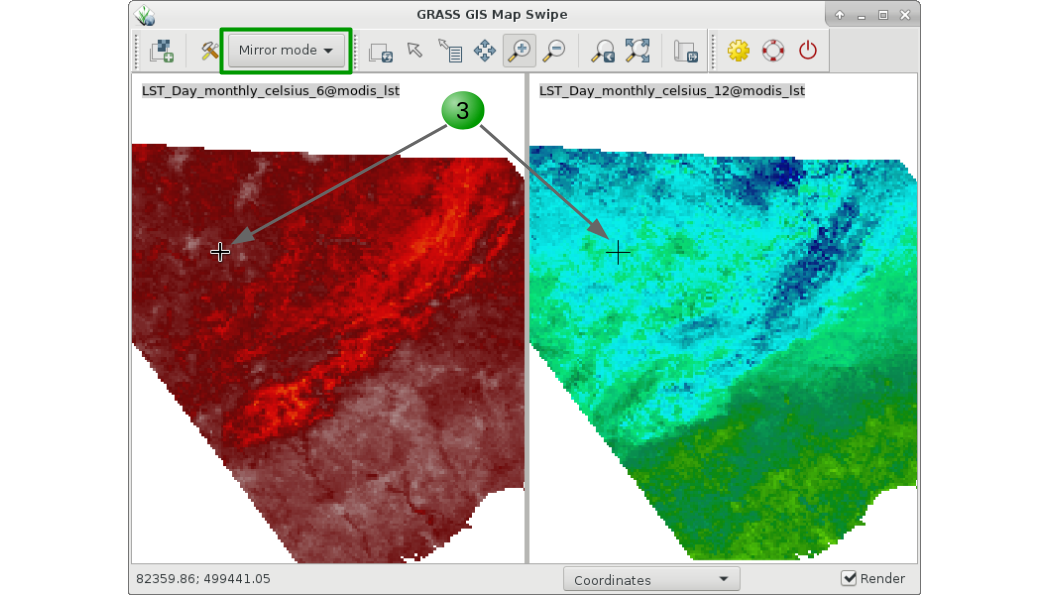
Aside from g.gui.timeline that we saw before, GRASS GIS offers different options to visualize time series data. The first one is g.gui.mapswipe that allows us to compare two maps from different dates for example by swiping a visibility bar. The module can be called from command line by specifying the two maps to compare or executed from the GUI as shown below in the respective tabs.
g.gui.mapswipe first=LST_Day_monthly_celsius_6 second=LST_Day_monthly_celsius_12
1. Select first and second map from the dropdown map lists.
2. Swipe as desired. Optionally you can change the swipe orientation or switch maps with the tools button.
3. Change to "Mirror mode" to see the exact same position in both maps.
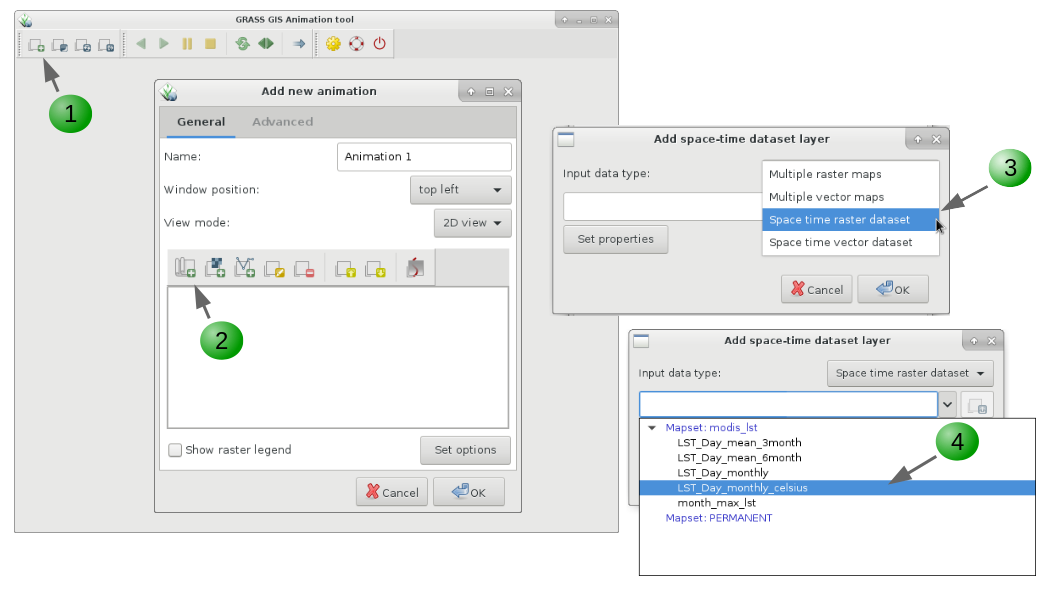
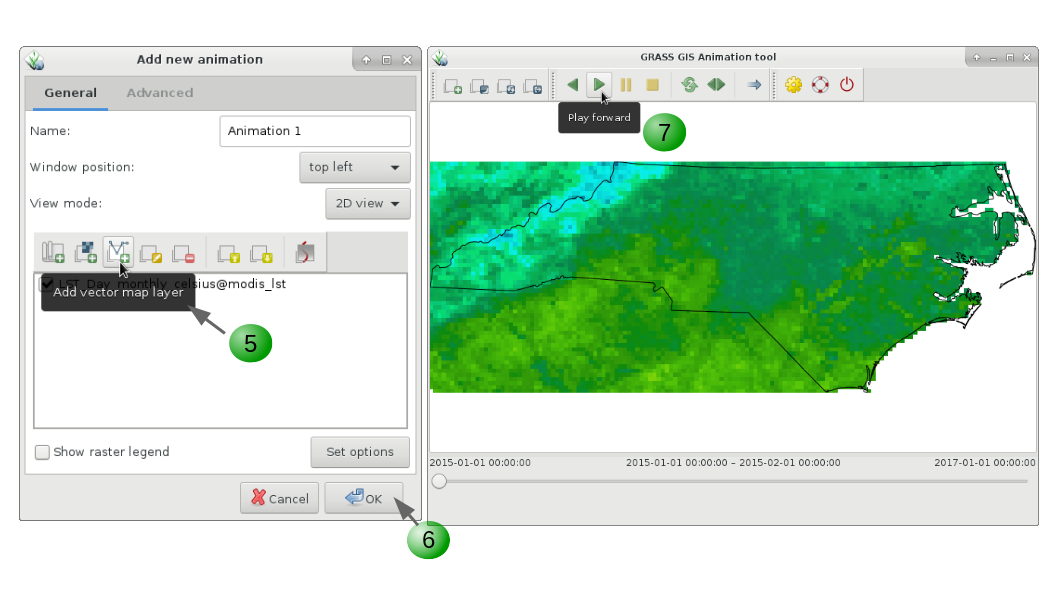
Another tool to visualize time series in more dynamic way and that allows us to see changes in space an time simultaneously is g.gui.animation. It also allows to animate vector time series, as well as lists of raster or vector maps not specifically registered as time series. When executed from command line, options are more limited than when the module is run from the GUI, where it is possible to add different decorations and customize several options.
g.gui.animation strds=LST_Day_monthly_celsius
1. Add a new animation (you can add up to 4 animations).
2, 3, 4. Add a space time data set (vector or raster) or list of raster or vector maps.
5. Optionally, add vector maps such as boundaries, points of interest, etc.
6. Once you finished adding maps and customizing their display, click OK.
7. Play the animation (note that you can move forwards or backwards, pause, stop, change velocity).
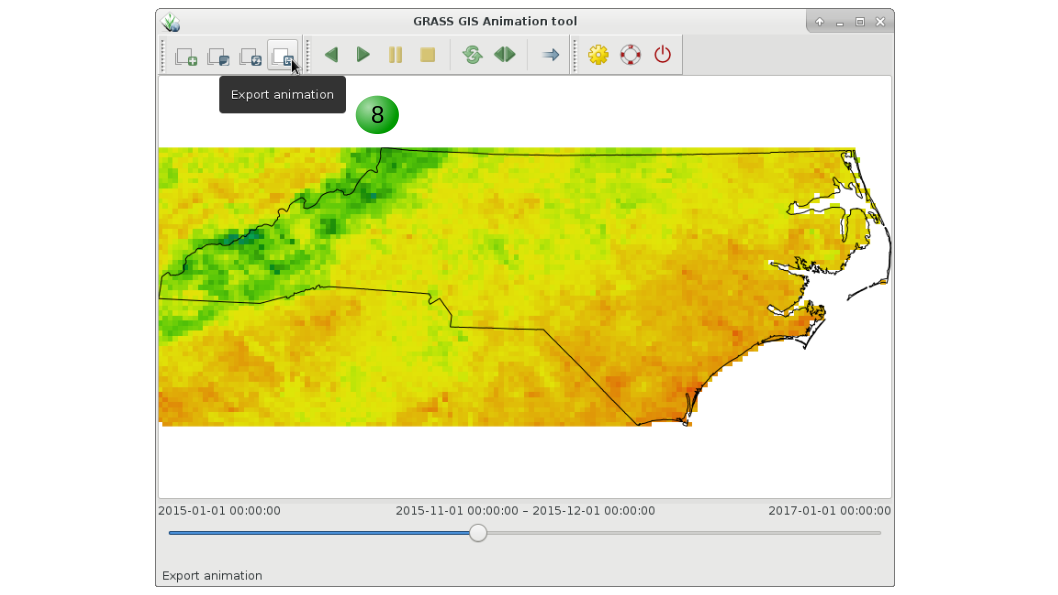
8. Export the animation.
9. Add decorations such as text, time stamp, images, if desired.
10. Select folder and file name (Note that this step slightly varies according to the output format desired).
11. Click Export.
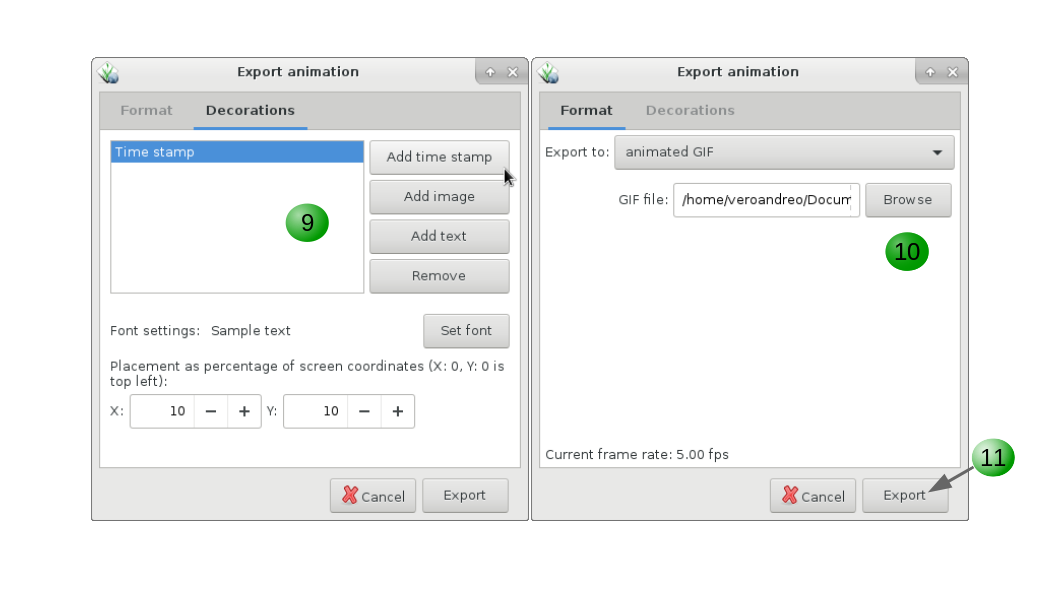
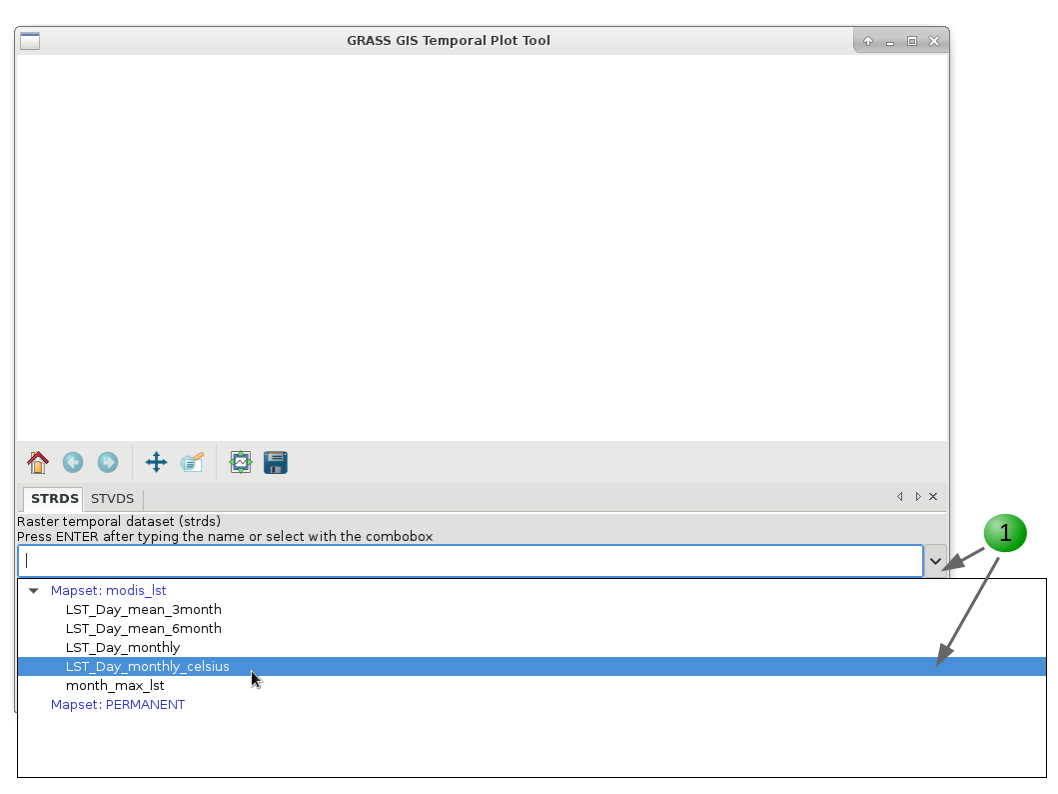
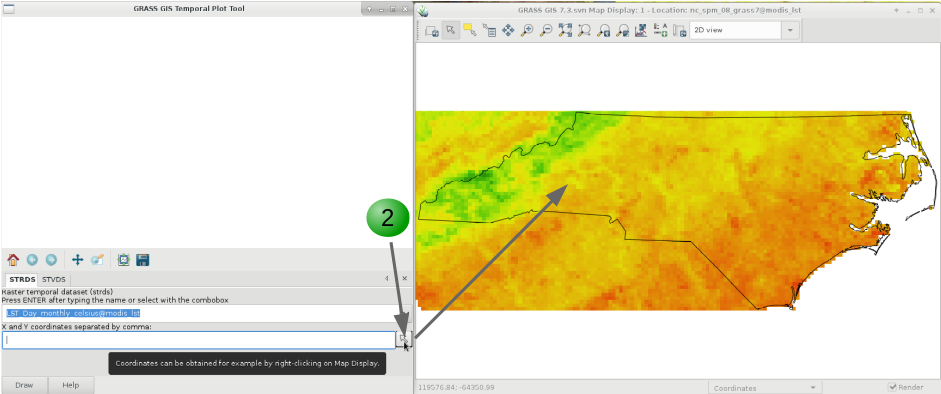
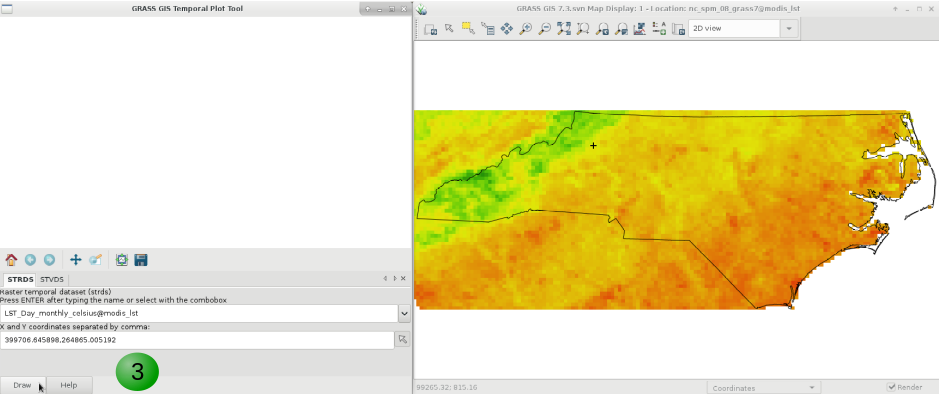
Finally, if you want to plot the time series of our variable of interest, be it in a STRDS or a STVDS, for a specific point of your study region, GRASS GIS has g.gui.tplot. Basically, you need to set the strds or stvds and a pair of X,Y coordinates. The latter can be typed directly, copied from the map display and pasted or directly chosen from the display. See instructions below.
g.gui.tplot strds=LST_Day_monthly_celsius coordinates=419604.018913,215736.80063
1. Select the STRDS.
2. Type coordinates or select a point in the display (use the arrow at the end of the second line).
3. Click Draw.
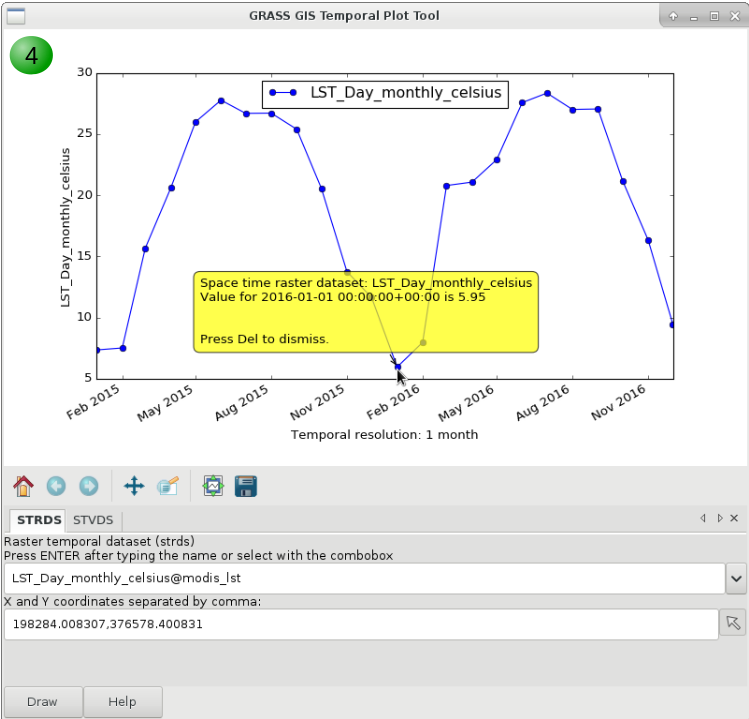
4. Inspect values for specific dates.
Alternatively, you can adjust the margin settings, zoom in to different parts of the plots and save it.
Sentinel satellite sensor data
GRASS GIS has a new set of extensions to manage Sentinel data. These tools allow to download, import, and preprocess Sentinel-2 data. Use g.extension to install them.
g.extension extension=i.sentinel
# install i.sentinel modules: i.sentinel.download, i.sentinel.import
# i.sentinel.preproc and i.sentinel.mask
g.extension extension=i.sentinel
gmod.Module("g.extension", extension="i.sentinel")
i.sentinel.download allows downloading Sentinel-2
products from Copernicus Open Access Hub.
To connect to Copernicus Open Access Hub a user name and password are required, see
Register new account page for signing up.
i.sentinel.download reads user credentials from a settings file.
Create the SETTING_SENTINEL file with the following content, e.g. in the directory $HOME/gisdata/:
myusername
mypassword
At this point you can use i.sentinel.download to search (using -l flag) and download Sentinel-2 scenes.
i.sentinel.download -l settings=$HOME/gisdata/SETTING_SENTINEL start=2017-07-30 end=2017-07-31 order=desc
i.sentinel.download settings=$HOME/gisdata/SETTING_SENTINEL start=2017-07-30 end=2017-07-31 order=desc out=$HOME/gisdata/ footprints=sentinel_2017_07
# search Sentinel-2 data for for the last days of July, 2017
# -l flag is used to print the resulting list
i.sentinel.download -l settings=$HOME/gisdata/SETTING_SENTINEL \
start=2017-07-30 end=2017-07-31 order=desc
# download data
i.sentinel.download settings=$HOME/gisdata/SETTING_SENTINEL \
start=2017-07-30 end=2017-07-31 order=desc out=$HOME/gisdata/ \
footprints=sentinel_2017_07
gmod.Module("i.sentinel.download", flags="l" settings="$HOME/gisdata/SETTING_SENTINEL",
start="2017-07-30", end="2017-07-31", order="desc")
gmod.Module("i.sentinel.download", settings="$HOME/gisdata/SETTING_SENTINEL", start="2017-07-30",
end="2017-07-31", order="desc", out="$HOME/gisdata/", footprints="sentinel_2017_07")
Once the data are downloaded you have to import them into GRASS GIS. There are two options: i.sentinel.import for a simpler import of the data, or i.sentinel.preproc to import and perform atmospheric correction.
i.sentinel.import requires only the input directory where the Sentinel-2 scenes were downloaded. Optionally, it is possible to select only some of the available bands. In the following example we are going to import only 7 bands for each image.
i.sentinel.import -rc input=$HOME/gisdata/ pattern='B(02|03|04|08|8A|11|12)'
# import the downloaded data
# -r flag is used to reproject the data during import
# pattern option is used to import only a subset of the available layers
# -c flag allows to import the cloud mask
i.sentinel.import -rc input=$HOME/gisdata/ pattern='B(02|03|04|08|8A|11|12)'
gmod.Module("i.sentinel.import", flags="rc", input="$HOME/gisdata/", pattern="'B(02|03|04|08|8A|11|12)'")
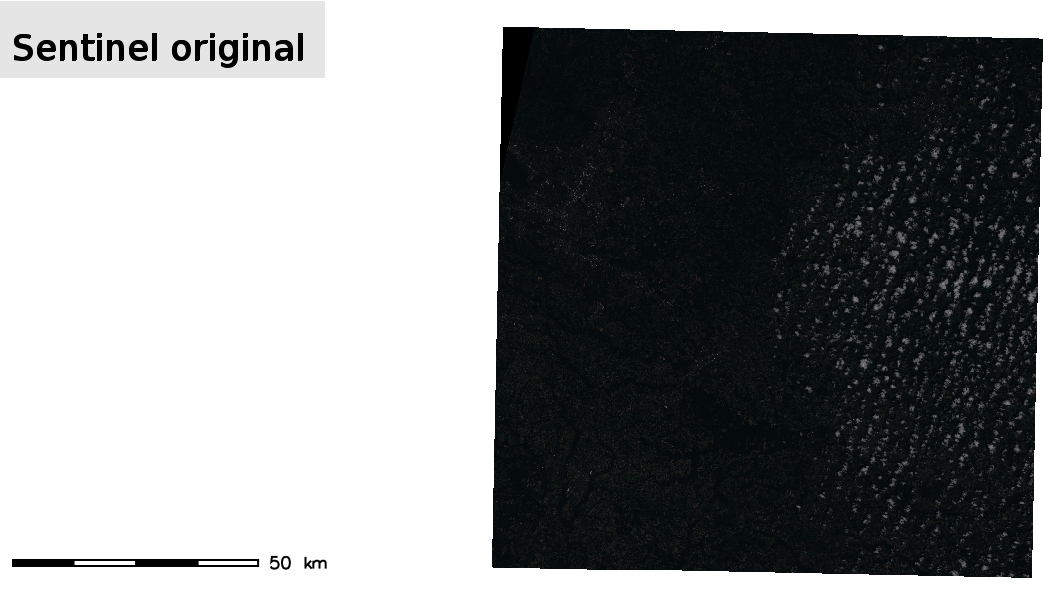
If we display the imported images, we can see that they appear really dark
d.mon wx0
d.rgb -n red=T17SQV_20170730T154909_B04 green=T17SQV_20170730T154909_B03 blue=T17SQV_20170730T154909_B02
d.barscale length=50 units=kilometers segment=4 fontsize=14
d.text -b text="Sentinel original" color=black bgcolor=229:229:229 align=cc font=sans size=8

To have a better visualization, it is required to perform color auto-balancing for RGB bands. The module to use is i.color.enhance. This module modifies the color table of each image band to provide a more natural color mixture, but the base data remains untouched.
i.colors.enhance red=T17SQV_20170730T154909_B04 green=T17SQV_20170730T154909_B03 blue=T17SQV_20170730T154909_B02
gmod.Module("i.colors.enhance", red="T17SQV_20170730T154909_B04", green="T17SQV_20170730T154909_B03", blue="T17SQV_20170730T154909_B02")
Now you can run again the previous piece of code to visualize the RGB combination of the Sentinel-2 scene and observe the difference.
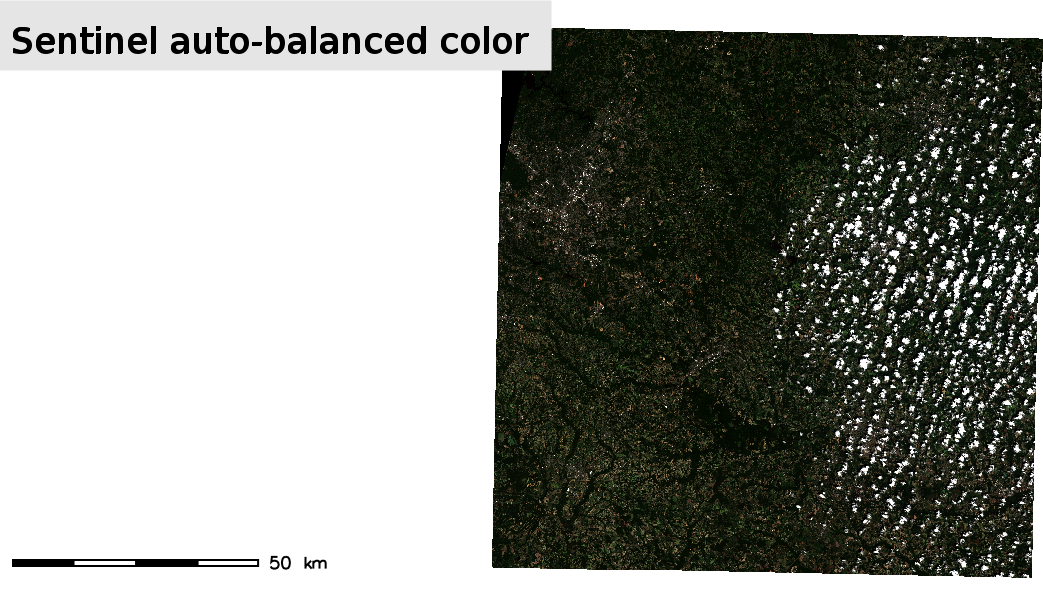
i.sentinel.preproc requires some extra inputs since it also performs atmospheric correction. First, this module requires the image as an unzipped directory, so you have to unzip one of the previous downloaded files, for example:
cd $HOME/gisdata/
unzip $HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.zip
Another required input is the visibility map. Since we don't have this kind of data, we will replace it with an estimated Aerosol Optical Depth (AOD) value.
It is possible to obtain AOD from http://aeronet.gsfc.nasa.gov. In this case, we will use the
EPA-Res_Triangle_Pk
station, select `01-07-2017` as start date and `30-08-2017` as end date, tick the box labelled as 'Combined file (all products without phase functions)'
near the bottom, choose 'All Points' under Data Format, and download and unzip the file into $HOME/gisdata/ folder
(the final file has a .dubovik extension).
The last input data required is the elevation map. Inside the North Carolina basic location there is an elevation map called elevation.
The extent of the elevation map is smaller than our Sentinel-2 image's extent, so if you will use this elevation map only a subset of the
Sentinel image will be atmospherically corrected; to get an elevation map for the entire area please read the next session.
At this point you can run i.sentinel.preproc
(please check which elevation map you want to use). The text_file option creates a text file useful as input for
i.sentinel.mask, the next step in the workflow.
i.sentinel.preproc -atr input_dir=$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE elevation=elevation aeronet_file=$HOME/gisdata/170701_170831_EPA-Res_Triangle_Pk.dubovik suffix=corr text_file=$HOME/gisdata/sentinel_mask
i.sentinel.preproc -atr input_dir=$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE \
elevation=elevation aeronet_file=$HOME/gisdata/170701_170831_EPA-Res_Triangle_Pk.dubovik suffix=corr text_file=$HOME/gisdata/sentinel_mask
gmod.Module("i.sentinel.preproc", flags="atr", input_dir="$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE",
elevation="elevation", aeronet_file="$HOME/gisdata/170701_170831_EPA-Res_Triangle_Pk.dubovik", suffix="corr", text_file="$HOME/gisdata/sentinel_mask")

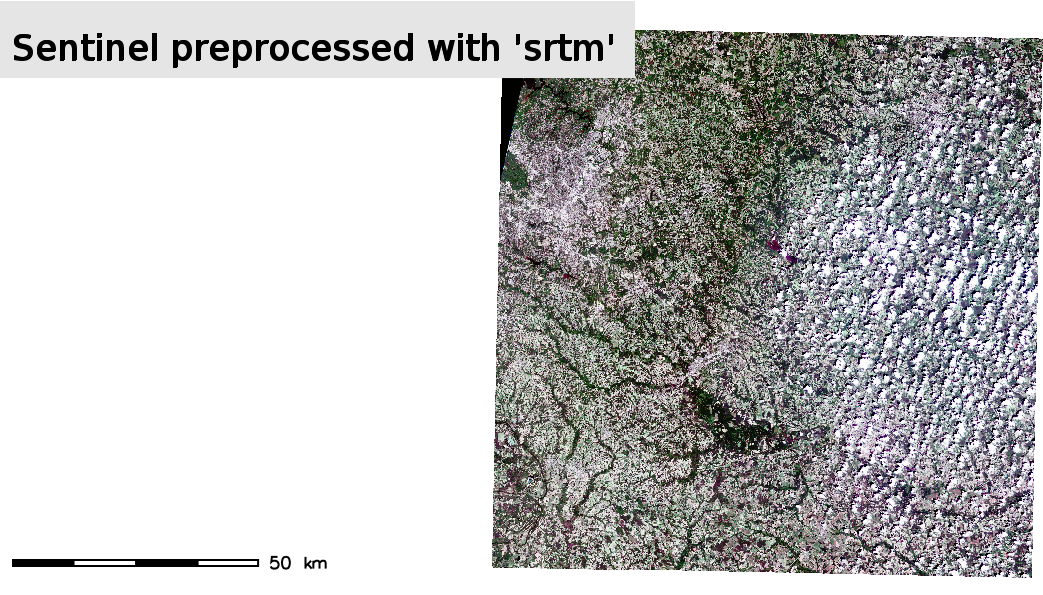
On the left the Sentinel-2 scene processed with elevation map, on the right the same scene processed with SRTM map
d.mon wx0
d.rgb -n red=T17SQV_20170730T154909_B04_corr green=T17SQV_20170730T154909_B03_corr blue=T17SQV_20170730T154909_B02_corr
d.barscale length=50 units=kilometers segment=4 fontsize=14
d.text -b text="Sentinel pre-processed scene" color=black bgcolor=229:229:229 align=cc font=sans size=8
Finally you can get the clouds and clouds' shadows masks for the Sentinel-2 scene using i.sentinel.mask.
i.sentinel.mask input_file=$HOME/gisdata/sentinel_mask cloud_mask=T17SQV_20170730T160022_cloud shadow_mask=T17SQV_20170730T160022_shadow mtd=$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE/MTD_MSIL1C.xml
i.sentinel.mask input_file=$HOME/gisdata/sentinel_mask cloud_mask=T17SQV_20170730T160022_cloud \
shadow_mask=T17SQV_20170730T160022_shadow mtd=$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE/MTD_MSIL1C.xml
gmod.Module("i.sentinel.mask", input_file="$HOME/gisdata/sentinel_mask", cloud_mask="T17SQV_20170730T160022_cloud",
shadow_mask="T17SQV_20170730T160022_shadow", mtd="$HOME/gisdata/S2B_MSIL1C_20170730T154909_N0205_R054_T17SQV_20170730T160022.SAFE/MTD_MSIL1C.xml")
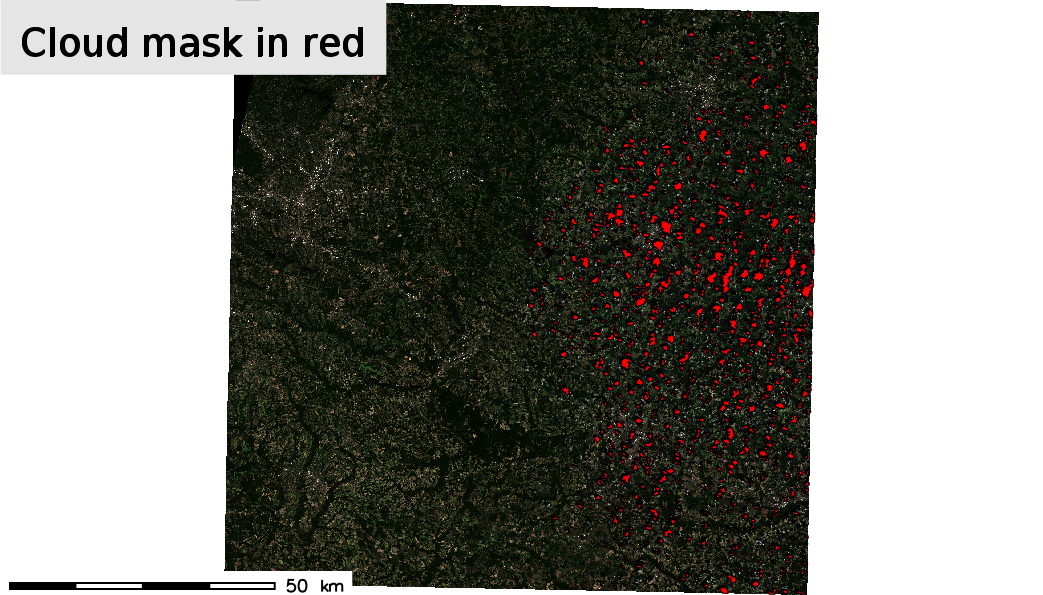
At this point we can visualize the output of i.sentinel.mask.
d.mon wx0
d.rgb -n red=T17SQV_20170730T154909_B04_corr green=T17SQV_20170730T154909_B03_corr blue=T17SQV_20170730T154909_B02_corr
d.vect T17SQV_20170730T160022_cloud fill_color=red
d.barscale length=50 units=kilometers segment=4 fontsize=14
d.text -b text="Cloud mask in red" color=black bgcolor=229:229:229 align=cc font=sans size=8
How to obtain SRTM digital elevation model
Shuttle Radar Topography Mission (SRTM) is a worldwide Digital Elevation Model with a resolution of 30 or 90 meters. GRASS GIS has two modules to work with SRTM data, r.in.srtm to import already downloaded SRTM data and, the add-on r.in.srtm.region which is able to download and import SRTM data for the current GRASS GIS computational region. However, r.in.srtm.region is working only in a Longitude-Latitude location.
First, we need to obtain the bounding box, in Longitude and Latitude on WGS84, of the Sentinel data we want to process
g.region raster=T17SQV_20170730T154909_B04,T17SPV_20170730T154909_B04 -b
g.region raster=T17SQV_20170730T154909_B04,T17SPV_20170730T154909_B04 -b
gmod.Module("g.region", flags="p", raster="T17SQV_20170730T154909_B04,T17SPV_20170730T154909_B04")
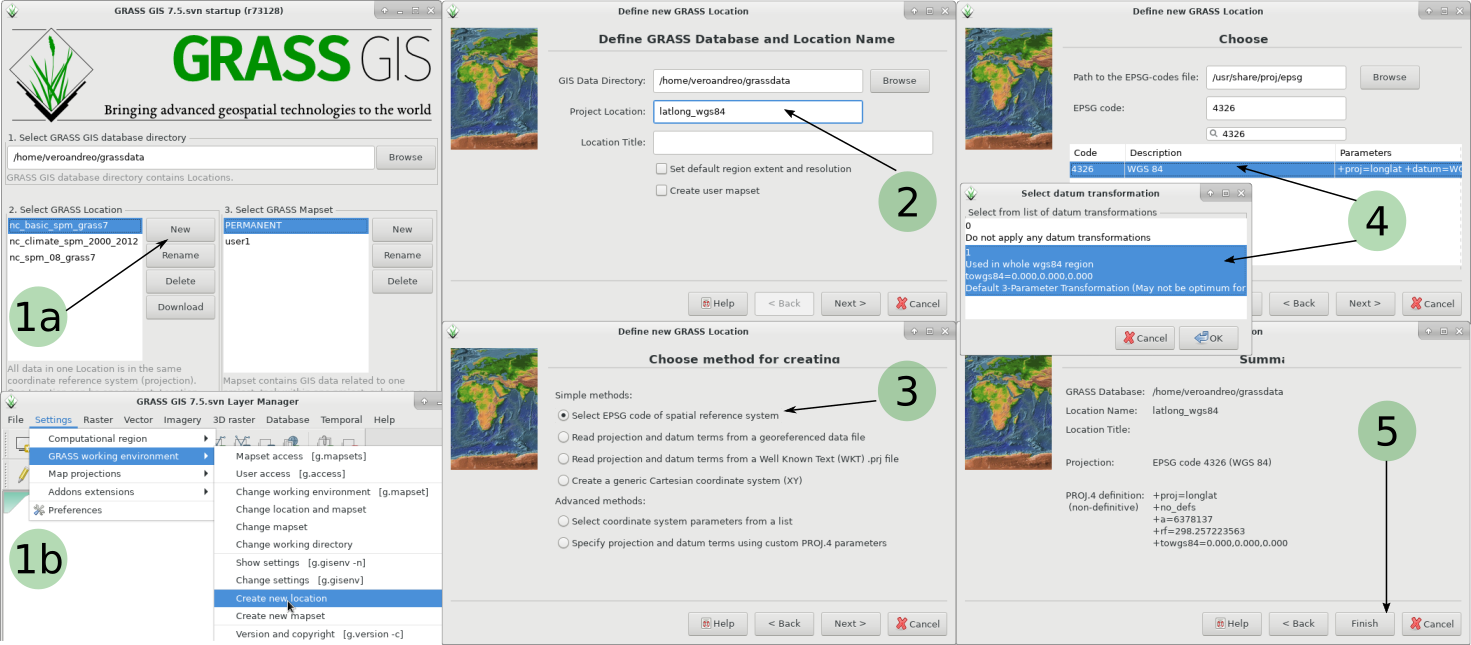
Now, we have to start a new GRASS GIS session in a Longitute-Latitude location
grass75 -c EPSG:4326 $HOME/grassdata/longlat1a. If in a new GRASS session, select "New" in the start-up screen 1b. Alternatively, from within a GRASS GIS session, go to "Settings --> GRASS Working environment --> Create new Location 2. Give a name to the new location 3. Select EPSG code as the option to create new location 4. Search for 4326 and select it 5. Done
from grass.script import create_location, gisenv create_location(gscript.gisenv()['GISDBASE'], "longlat", epsg=4326) gscript.run_command("g.mapset", location="mylonglat", mapset="PERMANENT")
Set the right region using the values obtain before
g.region n=36:08:35N s=35:06:24N e=77:33:33W w=79:54:47W -p
g.region n=36:08:35N s=35:06:24N e=77:33:33W w=79:54:47W -p
gmod.Module("g.region", flags="p", n="36:08:35N", s="35:06:24N", e="77:33:33W", w="79:54:47W")
After this you need to install r.in.srtm.region and run it
g.extension r.in.srtm.region
r.in.srtm.region output=srtm user=your_NASA_user pass=your_NASA_password
# install r.in.srtm.region
g.extension r.in.srtm.region
# run r.in.srtm.region downloading SRTM data and import them as srtm raster map
r.in.srtm.region output=srtm user=your_NASA_user pass=your_NASA_password
gmod.Module("g.extension", extension="r.in.srtm.region")
gmod.Module("r.in.srtm.region", output="srtm", user="your_NASA_user", pass="your_NASA_password")
You can now exit from this GRASS GIS session and restart to work in the previous one (where Sentinel data are).
To reproject the SRTM map from the longlat you have to use r.proj
r.proj location=longlat mapset=PERMANENT input=srtm resolution=30
r.proj location=longlat mapset=PERMANENT input=srtm resolution=30
gscript.run_command("g.mapset", location="nc_basic_spm_grass7", mapset="modis_lst")
gmod.Module("r.proj", location="longlat", mapset="PERMANENT", input="srtm", resolution=30)
At this point you can use srtm map as input of elevation option in i.sentinel.preproc
Other (very) useful links
- GRASS intro workshop: https://ncsu-osgeorel.github.io/grass-intro-workshop/
- Unleash the power of GRASS GIS: https://grasswiki.osgeo.org/wiki/Unleash_the_power_of_GRASS_GIS_at_US-IALE_2017
- Temporal data processing wiki: https://grasswiki.osgeo.org/wiki/Temporal_data_processing
- GRASS GIS temporal workshop: http://ncsu-geoforall-lab.github.io/grass-temporal-workshop/
- GRASS GIS and R for time series processing: https://grasswiki.osgeo.org/wiki/Temporal_data_processing/GRASS_R_raster_time_series_processing
- GRASS GIS Workshop in Jena: http://training.gismentors.eu/grass-gis-workshop-jena-2018/
References
- Gebbert, S., Pebesma, E. (2014). A temporal GIS for field based environmental modeling. Environmental Modelling & Software, 53, 1–12. DOI
- Gebbert, S., Pebesma, E. (2017). The GRASS GIS temporal framework. International Journal of Geographical Information Science 31, 1273-1292. DOI
- Neteler, M., Bowman, M.H., Landa, M. and Metz, M. (2012): GRASS GIS: a multi-purpose Open Source GIS. Environmental Modelling & Software, 31: 124-130 DOI
- Neteler, M., Mitasova, H. (2008): Open Source GIS: A GRASS GIS Approach. Third edition. ed. Springer, New York. Book site
Last changed: 2018-08-20 15:14
GRASS GIS manual main index | Temporal modules index | Topics index | Keywords Index | Full index

Big geodata management and analysis using GRASS GIS Workshop at FOSS4G 2018 by Veronica Andreo, Marco Ciolli, Luca Delucchi and Markus Neteler is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License - Thanks to Vaclav Petras for the style.